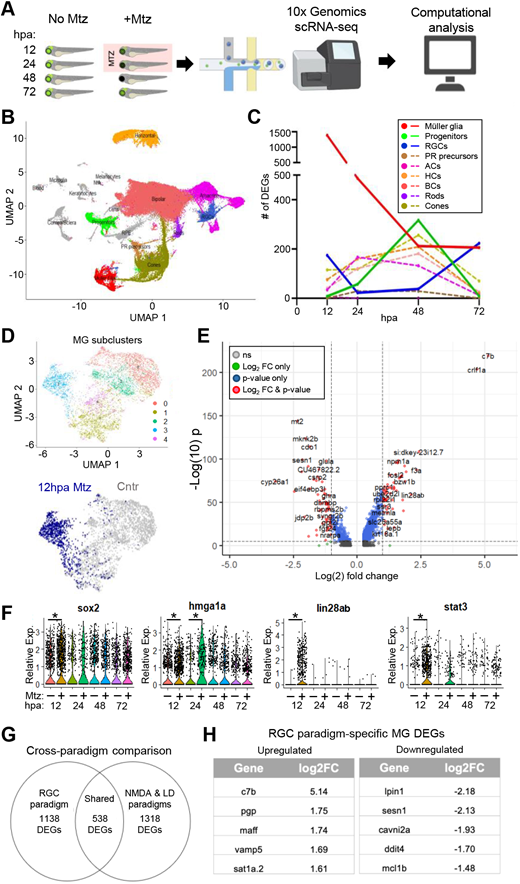
Fig. 2 Single cell transcriptomics of response to RGC ablation. (A) Experimental setup for scRNA-seq time series (12, 24, 48 and 72 hpa) of unablated control (no Mtz) and RGC-ablated (+Mtz) larvae. (B) UMAP of major eye cell types from all timepoints collected. (C) Total number of DEGs identified at each timepoint for MG, progenitor cells, RGCs and other major retinal cell classes. (D) MG subclusters. Subcluster 3 is made up predominantly of ‘activated’ MG cells at 12 hpa (blue). (E) Volcano plot of DEGs in 12 hpa MG subcluster. (F) Select MG DEGs at 12, 24, 48 and 72 hpa. (G) Cross-paradigm comparison of MG DEGs from RGC ablation data, and a pooled NDMA and LD dataset. (H) Top upregulated and downregulated RGC paradigm-specific DEGs relative to pooled NDMA and LD dataset. ACs, amacrine cells; BCs, bipolar cells; HCs, horizontal cells; PR, photoreceptor; RGCs, retinal ganglion cells. *P≤0.05.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Development