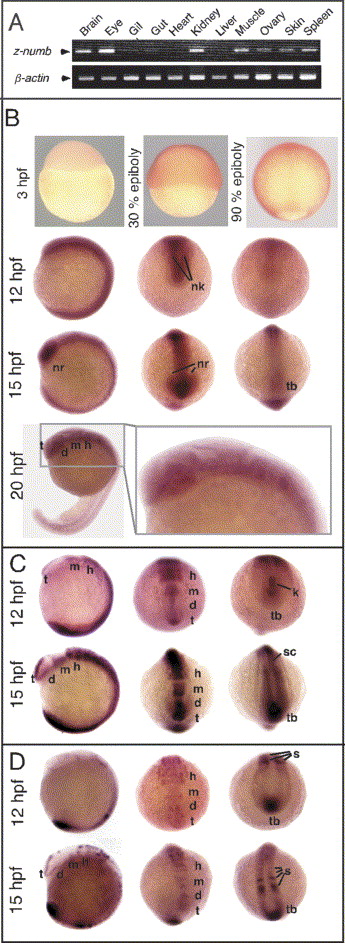
Fig. 2 Expression of znumb, notch1a, deltaD in zebrafish development. (A) Semi-quantitative RT-PCR analysis of znumb mRNA expression. Total RNA was isolated from various tissues of adult zebrafish, and znumb transcripts were amplified by using specific primers. μ-actin was used as a control. (B) Whole-mount in situ hybridization patterns of znumb expression during zebrafish embryonic development. Zebrafish embryos at various stages as indicated were hybridized with the antisense probe. Upper (3 hpf) panels are lateral views with the animal pole at the top. For the second (12 hpf) and third (15 hpf) rows, left panels are lateral view of embryos with anterior to the left, middle panels are frontal view with dorsal at the top, and right panels are a view of embryos from the posterior with dorsal at the top. A panel of 20 hpf in the fourth row is a lateral view with anterior to the left. Enlarged image is shown to the right side. (C,D) Whole-mount in situ hybridization patterns of notch1a (C) and deltaD (D). Left panels are lateral view of embryos with anterior to the left, middle panels are frontal view with dorsal at the top, and right panels are views from caudal to anterior with dorsal at the top. nk, neural keel; t, telencephalon; d, diencephalon; m, midbrain; h, hindbrain; kv, Kupffer′s vesicle; s, somites; tb, tail bud; nr, neural retina; sc, spinal cord. Sense probe hybridization was done for all samples, and no significant signals were observed (data not shown).
Reprinted from Mechanisms of Development, 123(5), Niikura, Y., Tabata, Y., Tajima, A., Inoue, I., Arai, K.I., and Watanabe, S., Zebrafish Numb homologue: Phylogenetic evolution and involvement in regulation of left-right asymmetry, 407-414, Copyright (2006) with permission from Elsevier. Full text @ Mech. Dev.