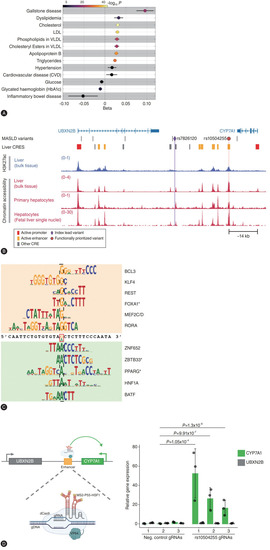
The association of relevant traits with the rs7826120 T allele (UK Biobank). (A) Associations were examined by additive linear or logistic regression analyses adjusted for BMI, age, sex, age×sex, age2 and age2×sex, first 10 genomic principal components and array batch. For binary traits, log odds of effects are shown. The colour represents the –log10-transformed P-values. (B) Overview of the metabolic- dysfunction associated steatotic liver disease (MASLD) locus UBXN2B/CYP7A1. Of all MASLD variants in the fine-mapped credible set at this locus, only rs10504255 (red circle) resides within an active liver cis-regulatory element (CRE). The lead variant (purple diamond) resides adjacent to an open chromatin region not enriched for the active CRE histone mark H3K27ac. Tracks show pooled normalised signal for H3K27ac chromatin immunoprecipitation sequencing (ChIP-seq) and chromatin accessibility detected by ATAC-seq across 4–6 independent samples. (C) MotifbreakR results showing position weight matrices (PWMs) of transcription factor (TF) motifs predicted to be affected by the variant rs10504255. PWMs matching the G allele, associated with higher proton density fat fraction, are shown at the top, and those matching the A allele are shown at the bottom. TFs with P<1E-3 were prioritised as likely hits if they were expressed in both liver tissue and hepatocytes and are shown ranked top to bottom for each allele by significance. *Indicates TFs for which there is ChIP-seq evidence of binding at the enhancer element. (D) Schematic (left) of CRISPRa complex targeting the enhancer at the UBXN2B/CYP7A1 locus. Bar plot (right) of relative expression detected by RT-qPCR of CYP7A1 and UBXN2B upon CRISPRa targeting of the enhancer vs. negative control guides. Data are presented as the mean±standard deviation of biological replicates (n=3 independent transductions) and statistically analysed by Student’s t-test.
|

