Fig. 5
- ID
- ZDB-FIG-230501-164
- Publication
- Jiang et al., 2022 - Parental mutations influence wild-type offspring via transcriptional adaptation
- Other Figures
- All Figure Page
- Back to All Figure Page
|
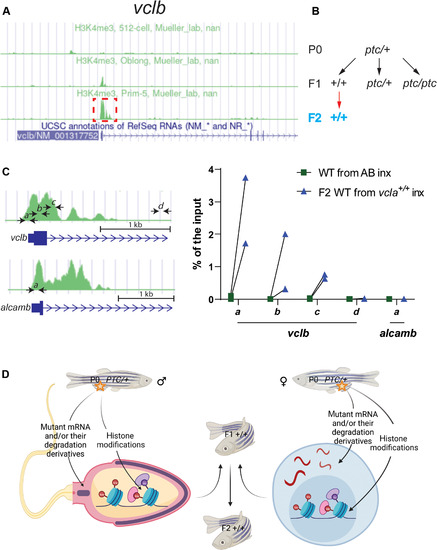
(A) Peaks of H3K4me3 ChIP-seq at the vclb locus at different stages (obtained from DANIO-CODE). Red boxed area was enlarged in (C) to show the relative location of the ChIP–qPCR primers marked by black arrows (also at the alcamb locus, which is used as an external/specificity control). (B) Genetic crosses were used to obtain F2 wild-type offspring from heterozygous zebrafish. Black arrows indicate heterozygous intercrosses (intx); red arrow indicates subsequent wild-type incrosses (inx). (C) ChIP-qPCR analysis of H3K4me3 occupancy near exon 1 of vclb in 512 cell stage wild-type embryos from incrosses of F1 vcla+/+ (i.e., wild types from vclaptc/+ intercrosses) compared with wild-type embryos from AB incrosses. Green peaks in vclb and alcamb loci represent prim-5 stage (24 hpf) H3K4me3 ChIP-seq data obtained from DANIO-CODE. Scale bars, 1 kb. n = 2 biologically independent samples. Ct values are listed in table S1. (D) Proposed model of intergenerational and transgenerational inheritance of TA (IGTA and TGTA). Mutant mRNAs, their degradation products (or their derivatives), and/or histone modifications are transmitted through the germline, leading to TA in the offspring. Additional mechanisms are likely to be involved in IGTA and TGTA. PTC, premature termination codon. (D) was created with BioRender.com. |