Fig. 1
- ID
- ZDB-FIG-211116-20
- Publication
- Black et al., 2020 - FOXM1 nuclear transcription factor translocates into mitochondria and inhibits oxidative phosphorylation
- Other Figures
- All Figure Page
- Back to All Figure Page
|
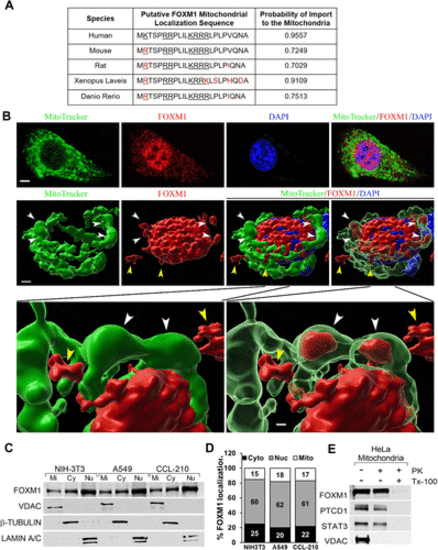
FOXM1 nuclear transcription factor is found in the mitochondria. (A) FOXM1 protein contains a highly conserved putative mitochondrial localization sequence (MLS). The MitoProt computational program predicted a positively charged (underlined) MLS in the N-terminus of FOXM1. FOXM1 MLS is highly conserved among several species (black font). Red color shows the differences compared with the MLS of human FOXM1. The probability of localization to the mitochondria for each FOXM1 orthologue was determined by MitoProt analysis. (B) Detection of FOXM1 in the mitochondria. High-resolution 3D confocal images show colocalization of FOXM1 (red) with mitochondrial marker MitoTracker (green) in NIH-3T3 cells. Nuclei were counterstained with DAPI. FOXM1 is present in the nucleus, mitochondria, and cytoplasm. White arrowheads point to the mitochondria with FOXM1 inside. Yellow arrowheads point to the cytoplasmic FOXM1. Magnification: ×100; 20 μm scale bar. (C) The presence of FOXM1 in mitochondrial (Mi), cytoplasmic (Cy), and nuclear (Nu) fractions is shown by Western blot with antibodies against FOXM1, VDAC (mitochondrial protein), β-TUBULIN (cytoplasmic protein), and LAMIN A/C (nuclear protein). (D) Quantification of FOXM1 protein levels in mitochondrial (Mi), cytoplasmic (Cy), and nuclear (Nu) fractions using densitometry. (E) Mitochondrial FOXM1 is proteinase K–resistant. Mitochondrial fraction was incubated without (lane 1) or with (lanes 2 and 3) 10 µg/ml proteinase K (PK). To disrupt mitochondrial integrity, 1% Triton X-100 (TX-100) was added in the digestion buffer (lane 3). Samples were probed for FOXM1, PTCD1 (mitochondrial matrix protein), STAT3 (inner membrane protein), and VDAC (mitochondrial outer membrane protein). |