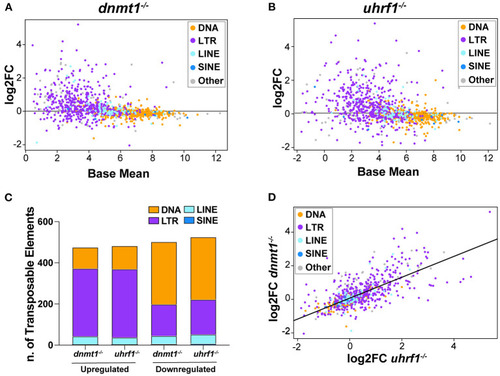
dnmt1 and uhrf1 loss causes overexpression of retrotransposons. RNAseq analysis of Transposable Elements in uhrf1−/− and dnmt1−/− mutant livers. (A) MA plot showing log2 fold change of repetitive elements in dnmt1−/− livers calculated on WT siblings and Base Mean in WT siblings. Repetitive elements are divided by Class in DNA transposons (yellow), LTR (purple), LINE (light blue), SINE (blue), and other (gray). (B) MA plot showing log2 fold change of repetitive elements in uhrf1−/− livers calculated on WT siblings and Base Mean in WT siblings. Repetitive elements are divided by Class in DNA transposons (yellow), LTR (purple), LINE (light blue), SINE (blue), and other (gray). (C) Bar graph of Transposable Elements divided by class in uhrf1−/− and dnmt1−/− mutant livers. Upregulated TEs have log2 fold change > 0 and downregulated TEs have log2 fold change < 0. (D) Correlation plot of repetitive elements in uhrf1−/− and dnmt1−/− mutant livers. Upregulated TEs have adj as pedix < 0.05 and log2 fold change > 0; downregulated TEs have padj < 0.05 and log2 fold change < 0. Log2 fold change is calculated between mutants and their own WT siblings.
|