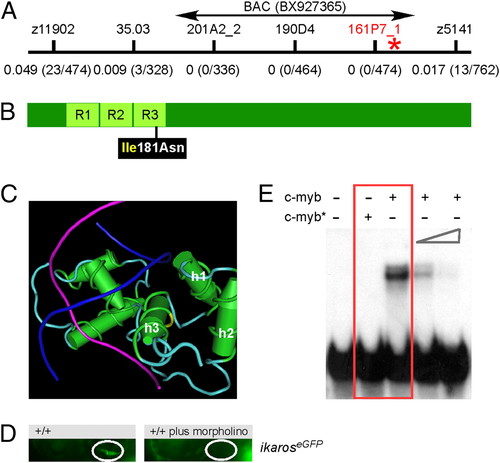
Fig. 2
|
A deleterious missense mutation in the DNA binding domain of zebrafish c-myb. (A) Genetic mapping of the IP109 mutation on chromosome 23. The locations of informative markers are indicated; recombination frequencies are given in brackets below their names (details in Table S3). The position of BAC BX927365 encompassing the c-myb gene that was used for complementation analysis is shown. (B) Schematic indicating the location of the isoleucine (Ile) to asparagine (Asn) missense mutation relative to the three repeat domains (R) of the DNA binding domain in the c-myb protein. (C) The side chain of isoleucine 181 in the third helix (h3) of repeat 3 (yellow area on the carbon trace) points away from the DNA double helix (strands in blue and magenta) to the first (h1) and second (h2) helix of repeat 3. This figure was rendered using Cn3D from PDB ID 1MSE (17). (D) Injection of an antisense c-myb morpholino oligonucleotide into wild-type fish transgenic for an ikaros:eGFP reporter recapitulates the thymic homing defect observed in c-myb mutants. The thymic rudiment is encircled. Lateral views, 4 dpf. (E) The mutant version of c-myb lacks in vitro DNA binding activity. The N-terminal one-half (amino acids 1–318) of the mouse c-myb protein, encompassing the highly conserved DNA binding domain, was expressed in E. coli; the I181N mutation was introduced by site-directed mutagenesis. The mutant protein (c-myb*) does not interact with the radioactively labeled DNA probe containing a c-myb consensus binding site (red box); excess unlabeled binding sites compete with the radiolabeled probe (two right-most lanes). Equal amounts of wild-type and mutant proteins were used (Fig. S2). |
| Fish: | |
|---|---|
| Knockdown Reagent: | |
| Observed In: | |
| Stage: | Day 4 |