1.0 INTRODUCTION
The technology of the World Wide Web (WWW) provides a revolutionary means for dissemination of scientific information. For the first time, scientists have 24 hour, low-cost international access to central repositories of research data without the need for specialized client-side software. In particular, biological researchers have exploited the power of the Web to create a diverse range of bioscience resources, including several web-accessible relational databases (e.g., Mouse Genome Database [8], the Human Genome Database [9], the C. Elegans database [5], the Genome Sequence Database [11], and FlyBase [7]). There is also a growing number of other Web sites serving static HTML documents; by spring of 1996 there were 26 different Web sites for 15 different species, and the number of sites is increasing at a rate of about one every three to four months.The biologists and computer scientists constructing these web sites presume that these web-accessible resources will aid scientific discovery through more timely, widespread access to better integrated research information. Although the WWW has made this information physically accessible to scientists, it is unclear whether it will be cognitively accessible. Busy scientists want useful, accurate, complete and up-to-date information without needing to learn and use a complex user interface. Will they be able to find answers to their questions without resorting to powerful but complex query languages like SQL? Will they be able to get their answers quickly without working through endless hierarchies of useless pages? Accessibility depends upon usability, and usability is critically related to productivity [10].
Designing a usable interface is challenging. First, the interface design must observe sound principles of graphic arts and psychology; it must have a functional layout, recognizable icons, and consistent interaction styles. Good design will reduce the time required to access desired information by minimizing misconceptions, mistakes and confusion in the search process. Second, it must incorporate a deep understanding of what information the scientist needs in the immediate context of his or her tasks and activities, and must present this information using language and conceptual models understood by the scientist.
We have created a Web-based biological database for the zebrafish research community. The success of our project and the achievement of true accessibility and productivity depend upon designing, developing, and implementing with a participatory design approach. The general merits of user-centered design methodologies have been widely discussed in the hypertext literature [15, 13, 17]. We have found, however, that the ubiquitous diversity in domain models, information access tasks, and experimental practices inherent to global scientific communities requires a more sensitive, participatory approach in which users are involved in every step of the design process. Here, we describe our use of participatory design [18, 6] and the unique challenges raised in applying this paradigm to a widely-distributed population of research scientists. Our experience demonstrates that, by using the WWW as a central, globally-accessible forum for design, participatory design techniques can be applied effectively to diverse, widely-distributed user communities.
2.0 OVERVIEW OF THE ZEBRAFISH INFORMATION NETWORK (ZFIN) PROJECT
Researchers using zebrafish to study basic biology, like genetics and development, are distributed among more than 100 laboratories in 28 countries. The zebrafish database project evolved out of our earlier Web site [19], which makes available (in static HTML documents) information on researchers and labs, a bibliography of publications relevant to the zebrafish research community, photos illustrating zebrafish developmental stages, and descriptions of laboratory methods, mutant lines, and the genetic map. The home page was accessed over 35,000 times in the past 18 months; in May 1997 alone, 14,000 HTML pages were served to 2600 sites located in 50 countries.Due to the exponential increase in information in this research area and the resulting need for more powerful methods of organizing and accessing these data, the zebrafish research community mandated extension of the original Web server to create a WWW accessible multimedia relational database known as the Zebrafish Information Network (ZFIN).
3.0 PARTICIPATORY DESIGN
Good design focuses on the ultimate usefulness and usability of an interactive software product by assessing the requirements and specifications of the product from the user's point of view, the user's interactive behavior when using the software, and the context of its use. We have found that the only way to realize this vision is to relocate the entire design process to the user's domain. Rather than integrating biological expertise into a traditional software design effort, our aim has been to integrate computer support into the everyday situated work activities of research geneticists. Achieving this goal required the software design team to essentially become adjunct members of a genetics research lab in order to understand the overall scientific process, the role of information access in that process, and the relationships that exist between individual researchers, laboratory groups, and the research community as a whole.We began by forging a design team which included both biologists and computer scientists. We believe that this close collaboration is key to the success of our project. Creating an abstract data model for the domain, formally articulating the research processes that users are engaged in, and structuring of interface actions to match information seeking tasks all require both biological and technical expertise to achieve efficiency and usability.
Designing for a Distributed Scientific Community
Applying the participatory methodology to a widely-distributed scientific community introduces a number of unique challenges. We identify three dimensions of difficulty: the heterogeneous nature of a scientific community, the lack of direct access to a significant portion of the community, and the technical challenges of interface design in the WWW environment. In this paper, we focus attention on the first two categories; technical challenges and our solutions to them are discussed elsewhere [3].To date, most participatory design efforts have been undertaken within large companies where the target users are (teams of) workers performing specific tasks within a tight collaborative framework defined by the company's overall production process. In contrast, a scientific community consists of loosely connected groups of relatively independent knowledge workers [2], each working on a particular aspect of the same general problem. Table 1 summarizes the special challenges encountered in designing for a scientific communities, contrasting them to corresponding features of a typical corporate context:
| Domain knowledge is extensive, diverse and very difficult to acquire, e.g., requires in-depth training in developmental genetics. The entities, processes, and relationships that define the basic structure of domain knowledge (i.e. domain ontology) are constantly changing as the research discipline evolves; these changes are not instantaneous, taking months or years to propagate through the community. | Domain knowledge may be extensive and fast-growing. However, the basic business model and the relevant domain entities and procedures associated with it are usually established by executive decree and are relatively stable. | |
| Many forms of formal and informal information exchange exist. Formal information exchange (e.g. publication) is highly institutionalized to guarantee accuracy and proper attribution; informal information exchanges abound and are ever-changing as workers migrate between labs, people enter and leave the community, and projects change. | Well-defined channels of information exchange have been defined by management. Informal means of information exchange exist as well, but are generally stable once established. | |
| Labs form the basic social group, with each lab headed by a principal scientist and containing other research scientists, post-docs and doctoral students. Labs are loosely-connected into a global scientific community. Although certain scientific standards exist, there is much variation in the detailed research practices (e.g. measurement, data collection) of different labs. | Work groups form the basic social group, forming the leaves of a hierarchical corporate structure. A uniform set of working procedures for each group is dictated from above; work groups are ultimately directed by a single executive entity. | |
| Labs and individual scientists within them are both cooperative and competitive. Progress of the discipline depends on sharing research results; success of the individual depends on attribution of results to the individual. | Success of the individual is intimately tied to the success of the work group and, more generally, to the success of the corporation. Unfettered information flow within the company is encouraged. |
In general, Table 1 emphasizes the independent, heterogeneous nature of scientific communities. This has forced us to modify the participatory design process in a number of ways. Rather than studying and designing for a single, representative group of users, we have had to come to grips with the fact that no such group exists in a diverse scientific community. Although, from a practical perspective, intensive ethnographic user analysis must be reserved for one or two highly-accessible groups of users, we have invested much effort in sharing the resulting insights with the remainder of the community, generalizing our formal models of domain activities and data to encompass the differences that exist within the community.
A simple example is the representation of developmental time within our abstract data model. Our initial analysis characterized the developmental age of an embryo in terms of a closed set of named developmental "stages" defined by the zebrafish community. However, subsequent testing of this characterization against the work practices of the broader research community revealed that different laboratories have different conventions for specifying developmental age, e.g., recording the age of embryos in hours or, more coarsely, in days. The data model had to be generalized to accommodate these different metrics while still supporting efficient indexing and retrieval of data.
A related difficulty is the continuously evolving conceptions of what entities, processes, and relationships exist or are relevant in a scientific domain. In most corporate contexts, this abstract model of the domain, also known as the domain ontology, is relatively stable. Although new information may accrue very rapidly, the kinds of information of interest generally remain the same or change only very slowly. For example, the domain model for a bank includes conceptual entities like accounts, balances, credit-to-debt ratios, and so on; this model has remained relatively stable for decades. This is not true in scientific domains, where the domain ontology is continually being modified and extended as the science evolves, new experimental techniques produce new kinds of data, and new biological entities are distinguished. Thus, we had to not only allow for easy changes and extensions to the domain data model, but also to support explicitly the gradual evolution of the data model from one state to the next without disrupting performance.
Perhaps the most confounding factor in designing shared data resources for scientific communities is the tension between information sharing and secrecy. Because research geneticists are all essentially working on the same problem (i.e., connecting genetic code with biological characteristics) and because experiments are extremely time- and effort-intensive, sharing of information as soon as it becomes available is highly desirable. At the same time, the success of individual scientists hinges on publishing unique scientific results, motivating researchers to keep information to themselves. A viable solution to this dilemma must maintain proper accreditation of work while making new results available soon after they are discovered.
The widely-distributed nature of scientific communities interferes directly with one of the basic tenets of participatory design, namely the immersive involvement of designers in the everyday workings of the community. This limitation makes it particularly difficult to expose the widespread variations in domain knowledge structure and scientific practice that exist within the community, so that they can be supported in the design. Accordingly, a primary challenge was to find ways of adapting the participatory design process to a distributed design context, balancing the need for intensive ethnographic analysis with the need to expose and accommodate the diverse requirements of the entire community of users. As discussed in the following section, we addressed this difficulty using a two-pronged approach, capitalizing on face-to-face collaborative contact with community members whenever possible, while utilizing the WWW to disseminate information and collect feedback on the nascent data model and the information access tasks to be supported by the proposed scientific database.
Participatory Design: Process
The distinction between the participatory and user-centered design paradigm centers on the composition of the design team and the status accorded to users in the design process [18, 6]. Rather than seeing end-users as "clients" from whom requirements are extracted and against whom (eventually) prototypes are tested, participatory design accords end-users first-class membership in the design team, giving them an active role in every part of the design process. We have found that this tight collaboration helps both domain experts and software engineers to develop and maintain a deep, shared understanding of each other's perspectives on the evolving design. In addition, the intimate participation of end-users fosters a level of commitment to the design within the user community that we have never before encountered using other design paradigms. This commitment by high-status scientists is essential in promoting the acceptance of new technology by the rest of the community.The basic structure of the design process is the same for both participatory and user-centered approaches; our design process (Figure 1) follows the basic steps of user-centered design described in texts like [12]. In the following paragraphs, we describe our execution of each design phase, with particular emphasis on how we addressed the unique challenges associated with participatory design for a widely-distributed scientific community:
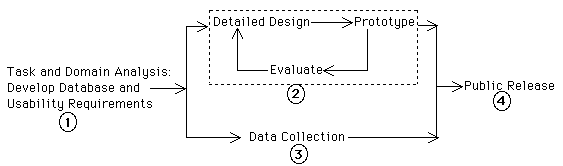
Step 1: Develop database and usability requirements
The initial step in our design process is to conduct domain and task analyses. The goal of this step is to produce both database information and interactive system specifications. However, the biological domain is extremely specialized, making this analysis very difficult. Development of the abstract data model, the nomenclature used to label user interface components, and the structuring of interface actions into information seeking tasks all require deep knowledge of the domain to achieve efficiency and usability. Accordingly, we began the design process by forging a participatory design team [18, 6] which includes both biologists and computer scientists. We believe that this collaboration is key to the success of our project, not only because it provides domain and task knowledge, but also because direct involvement of biologists gives them a stake in the success of the project.During Step 1 (Figure 1) we used primarily ethnographic methods [1], including interviews with zebrafish scientists, reading journal articles, attending research talks and lab meetings, and participant-observer activities such as helping to customize the specialized software used by one of the labs. In other words we tried to "go native" in the zebrafish community. We also used questionnaires to gather design input from scientists around the world, distributing them at workshops, and via the original Zebrafish Web site, which contains documents on the development of the database project along with a brief list of the types of information we expect to include. We solicited feedback from our users about their satisfaction with the current HTML-based Web site and integrated their requests for enhanced functionality and information into the design of the new database. The goal of this immersion in the working world of zebrafish scientists was to understand the context of their everyday work activities and its relationship to the proposed WWW database.
In addition to these ethnographic methods, we looked extensively at other web-accessible biological database sites to evaluate their information content and user interfaces; we videotaped our own zebrafish scientists doing simple information retrieval tasks to pinpoint their confusions with the user interfaces and information models at these other sites. In this respect, we have found the WWW to be an easily accessible means of drawing on the design experience of others, a critical resource in an area where design improvements are typically incremental and based on real-world experience.
These domain and task analyses produced specifications for the database information and the user interface. Database information specifications were captured in a data model document intelligible to both computer scientists and biologists, containing descriptions of database entities, attributes, and relationships, as well as examples of situations of use for various pieces of information. The data model serves as the blueprint for database implementation and offers zebrafish biologists a concise overview of database contents. The current design for the database is complex and incorporates 21 major classes of information, most of which are highly interconnected [20].
Step 1 (Figure 1) also produced specifications for the interactive system. In particular, functional and performance requirements of the system were determined as seen from the user's point of view. The first of these, functional requirements, describes what the system should do. Our functional requirements include:
- Must provide a security mechanism to ensure that only authorized users submit data.
- Must guarantee reliability and completeness of the data, especially because much of it is user supplied.
- Must have a mechanism to distinguish published data from unpublished or pre-published data.
- Must allow submission of commonly published image types, including annotations.
- Must provide color reliability and reasonable resolution for images so that information is accurate enough to make science-based decisions.
- Must be accessible from multiple platforms, including older machines, to support universal access to the database.
- May need to access other databases concurrently with ZFIN.
- Must be easy to learn. We expect user interactions with the database (retrievals or submissions) to be relatively infrequent, and thus, a typical scientist might forget how to use the interface between uses. The interface must be learned as you go (no separate manual) and must provide extensive on-line help.
- Must be fast enough to satisfy most commonly asked questions within a 10 minute session.
- Must provide enough feedback that most searches will find results with three rounds of querying.
- Must keep the user apprised of progress during the data submission process, allowing the user to "undo" a submission during any step.
Step 2: Iterate detailed design process
Step 2 (Figure 1) is the heart of usability engineering methods [14]. After developing the information and usability requirements, we moved into the iterative refinement phase of the participatory design process to design and implement the user interface. In contrast to the traditional waterfall approach, this technique relies on rapid cycles of design, prototype implementation, and evaluation with real users to generate the final product.Rather than implementing the entire database at once, we initially selected a subset of the database information (i.e. data types) to take through Steps 2-4 of the design process. This was done primarily for pragmatic reasons. Some information is of higher priority to the research community or more mature and complete; staggering development of various data classes allowed us to make useful information rapidly available. In addition, focusing attention on just a few types of information at a time proved to be an effective means of managing the complexity of the design.
Each prototype was immediately evaluated by selecting pairs of zebrafish scientists and giving them typical data retrieval and submission tasks to perform. Videotaping these sessions allowed us to analyze the amount of time required, misconceptions encountered, and other problems with the interface. We evaluated their performance against the usability requirements developed in Step 1 (Figure 1). Details of how to conduct this type of performance analysis can be found in [4]. We used insights gained from this analysis to shape subsequent prototypes in the iterative design cycle.
When we were satisfied with the usability of a prototype, it was made available via the WWW to 15 zebrafish scientists (acting as beta testers) distributed among both large and small labs around the world; access to the prototypes was limited to these testers. Feedback on the prototype was gathered using a series of short, on-line questionnaires and by including a "comment" button on each screen of the prototype, allowing users to easily generate an email message to the developers. In addition, the system recorded the screens traversed by each user, allowing us to identify areas of particular interest and to expose problems with the interface. At the end of the beta testing period, we also interviewed our testers by telephone to discover any other problems. The prototypes were also demonstrated at special sessions during professional meetings, with feedback gathered orally and by distributing short questionnaires.
As a result of this participatory process, we have developed a committed group of "co-developers" in the user community that is highly representative of the global community as a whole. As an unexpected benefit, we found that our participatory approach has greatly increased enthusiasm for the ZFIN project within the research community. For example, during the first weeks of the beta testing phase, our data editor was deluged with requests to be added to the "community members" portion of the database, as scientists learned (apparently from the beta testers) of the nascent data resource. We feel that this level of enthusiasm stems directly from our users' sense of involvement in the design process, and will play an important role in the ultimate success of the ZFIN database.
Steps 3 and 4: Data Collection and Public Release
Because data collection (Step 3) is a relatively mundane part of the design process, it will not be addressed extensively here, in favor of a more focused discussion of usability and interface design issues. Briefly, data editors were recruited to gather existing domain data from various sources (e.g. publications, lab notes, existing data archives). These data were then mapped to the data model developed in Step 2 to populate the database.Step 4, public release of the database, is an important part of the participatory design process rather than its abrupt termination; usability analysis will continue indefinitely, allowing the system to evolve to meet the changing needs of users. The commentary forms for gathering user feedback mentioned earlier in the context of beta testing remain available in the public release. We are also recording (anonymously) the sequence of screens visited by each user and the total number of visits to each screen to determine common usage patterns and to expose areas of confusion. Finally, we are planning to conduct extensive periodic user surveys, interviewing a sample of randomly selected registered ZFIN users to assess usage patterns, good and bad features of the Web site, and interest in future information support.
5.0 CONCLUSIONS
Rapidly expanding access to the WWW holds incredible promise for increased data sharing and collaboration within widely-distributed research communities. Web-accessible databases will make available a much broader range of data than printed media, including multimedia data types and information that, although useful, might never be formally published; new findings can be made available almost instantly, rather than being delayed for months by a lengthy editorial and printing process.There are a number of challenging obstacles to such universal accessibility. First, scientific domains are unusually complex, with domain models that evolve with the expanding frontiers of the discipline; the kinds of information accepted as "data" and the research techniques that generate this information change over time. Consequently, interface design for a scientific database is much more demanding than for stable, single-use databases.
Our experience with this project demonstrates that participatory design can be used to manage domain complexity and generate meaningful usability requirements by focusing attention on the real-world work activities (i.e. research processes) of users and on the ways in which access to various kinds of information (data) contributes to these activities. It also demonstrates that, by using the WWW as a medium for both the design process and the design itself, participatory techniques can be adapted to contexts in which the user population is widely-distributed and loosely-organized.
We have focused our work to date on maximizing the accessibility of data contained in the ZFIN database, applying participatory techniques to streamline individual data manipulations. In the future, we plan to expand our focus to support the full research process in which database access is embedded by developing tools to support database-centered interaction among widely-distributed members of the research community. Examples include mechanisms for collaborative shared access to the database, interactive discussion forums, and viewer commentary appended to specific data records. We envision ZFIN as the cornerstone of a virtual community of research scientists linked via the WWW, working together, and sharing a common set of data.
Acknowledgments
Mike McHorse and Paul Bloch provided essential system administration support for our computers and network. We would also like to thank our numerous informants within the zebrafish research community, who patiently took time to explain their research methods to us. The zebrafish database project is sponsored by the W.M. Keck Foundation and NSF grant BIR-9507401.References
- Blomberg, J., J. Giacomi, A. Mosher, and P. Swenton-Wall. (1993), Ethnographic field methods and their relation to design, in Participatory Design: Principles and Practices, D. Schuler and A. Namioka, Editors. Lawrence Erlbaum Associates: Hillsdale, NJ. p. 123-155.
- Crane, D. (1972), Invisible Colleges. Chicago: University of Chicago Press.
- Doerry, E., S.A. Douglas, A.E. Kirkpatrick, and M. Westerfield. (1997), Moving beyond HTML to Create a Multimedia Database with User-Centered Design: A Case Study of a Biological Database, University of Oregon Technical Report CIS-TR-97-02: Eugene, OR.
- Douglas, S.A. (1995), Conversation analysis and human-computer interaction design, in Social and Interactional Dimensions of Human-Computer Interfaces, P.J. Thomas, Editor. Cambridge University Press. p. 184-203.
- Genome Informatics Group (1996), ACEDB, US Department of Agricultur (World Wide Web URL http://probe.nalusda.gov:8300/cgi-bin/query?dbname=acedb).: Beltsville, MD.
- Greenbaum, J. and M. Kyng (1991), Design at work: Cooperative design of computer systems. Hillsdale, NJ: Lawrence Erlbaum Associates.
- Harvard Medical School (1996), FlyBase, (World Wide Web URL http://cbbridges.harvard.edu:7081/), Harvard Medical School: Cambridge, MA.
- Jackson Laboratory (1996), Mouse Genome Database, (World Wide Web URL http://www.informatics.jax.org/): Bar Harbor, Maine.
- Johns Hopkins School of Medicine (1996), Genome Database (GDB), (World Wide Web URL http://gdbwww.gdb.org/): Baltimore, MD.
- Landauer, T.K. (1995), The trouble with computers. Cambridge, MA: MIT Press.
- National Center for Genome Resources (1996), Genome Sequence Database (GSDB), (World Wide Wed URL http://www.ncgr.org/gsdb/): Santa Fe, NM.
- Newman, W.M. and M.G. Lamming (1995), Interactive systems design. Wokingham, England: Addison-Wesley.
- Nielsen, J. (1995), Multimedia and hypertext: The Internet and beyond. (Chap. 10, pp. 279-307). Boston: Academic Press
- Nielsen, J. (1993), Usability Engineering. Boston: Academic Press.
- Nielsen, J. (1989), Evaluating hypertext usability. In D. H. Jonassen and H. Mandl (Eds.), Designing Hypermedia for Learning, (pp. 147-168). New York: Springer-Verlag.
- Greene, S.L., S.L. Gomez, and S.J. Devlin (1986), A cognitive analysis of database query production. In Proceedings of the Human Factors Society. Santa Monica, CA: Human Factors Society.
- Perlman, G., D. Egan, K. Ehrlich, G. Marchionini, J. Nielsen, and B. Shneiderman. (1990), Evaluating hypermedia systems. In Proceedings of Human Factors in Computing CHI'90 Conference, (pp. 387-390). New York: ACM Press.
- Schuler, D. and A. Namioka (1993), Participatory design: Principles and practices. Hillsdale, NJ: Lawrence Erlbaum Associates.
- University of Oregon Institute of Neuroscience (1996), The FISH Net, (World Wide Web URL http://zfin.org): Eugene, OR.
- Westerfield, M., E. Doerry, A.E. Kirkpatrick, W. Driever, and S.A. Douglas. (1997), An on-line database for zebrafish development and genetics research. Sem. Devel. Biol. (in press).
