- Title
-
3DM: deep decomposition and deconvolution microscopy for rapid neural activity imaging
- Authors
- Cho, E.S., Han, S., Lee, K.H., Kim, C.H., Yoon, Y.G.
- Source
- Full text @ Opt. Express
|
Fig. 1. Deep decomposition deconvolution microscopy (3DM). (a) Schematic of 3DM hardware. An electrically tunable lens (ETL) is conjugated to the back pupil plane of the objective lens. (b) Schematic of the 3DM algorithm. A raw video consists of a time series of 3-D volumes. Using a bilinear neural network for efficient approximation of RPCA (BEAR), the raw video is decomposed into a low rank component and a sparse component that correspond to the background and the neural activity, respectively. The decomposed sparse component is deconvolved using our 3-D deconvolution network. |
|
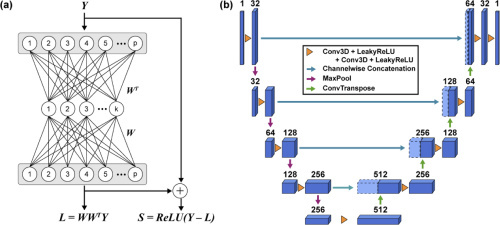
Fig. 2. Neural networks for sparse decomposition and deconvolution. (a) Architecture of BEAR. The network takes the raw video
|
|
Fig. 3. Temporal maximum intensity projection (MIP) of the sparse components of the simulated images. Scale bar, 100
|
|
Fig. 4. Imaging whole brain and spinal cord of a larval zebrafish expressing pan-neuronal GCaMP7a using 3DM. (a) Temporal MIP of the raw images. Scale bar, 100
|
|
Fig. 5. Imaging the neuronal activity of a larval zebrafish brain expressing pan-neuronal GCaMP7a using 3DM. (a) Three z-slices (at z=70
|