- Title
-
Non-acylated Wnts Can Promote Signaling
- Authors
- Speer, K.F., Sommer, A., Tajer, B., Mullins, M.C., Klein, P.S., Lemmon, M.A.
- Source
- Full text @ Cell Rep.
|
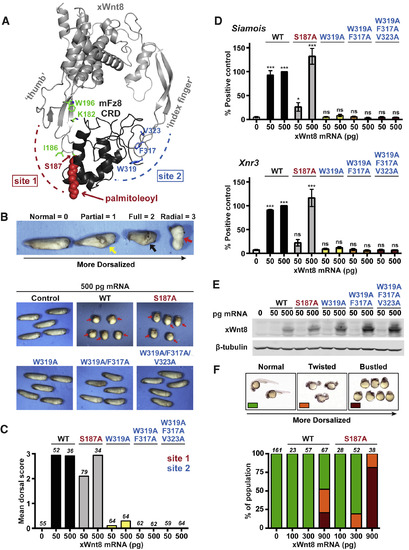
Effects of Site 1 and Site 2 Mutations on xWnt8 Activity (A) Crystal structure of the xWnt8/mFZD8 CRD complex (PDB: 4F0A), with “thumb” and “index finger” projections on xWnt8 binding to the CRD at sites 1 and 2, respectively (Janda et al., 2012). Residues mutated in site 1 (green) and site 2 (blue) are marked. The palmitoleoyl chain and S187 are red. (B) Dorsalization phenotypes observed upon ectopic xWnt8 expression in ventral cells of X. laevis embryos. The top panel shows tailbud-stage embryos with archetypal Wnt overexpression phenotypes and corresponding scores. Representative xWnt8 mutant phenotypes are shown in the bottom two rows. Yellow arrow, partial axis duplication; black, full axis duplication; red, radial dorsalization. (C) Quantitation of dorsalization phenotypes in Xenopus embryos for site 1 and site 2 mutations. Total number of embryos scored (across three biological replicates) is listed for each bar. Dorsal scores for xWnt8WT and xWnt8S187A are from the dataset in Figure 2A, represented here for comparison. (D) Initial RT-PCR quantitation of Siamois and Xnr3 induction for each variant, represented as mean ± SEM (n = 3). Significance is denoted as ns (p ≥ 0.05), ∗ (p ≤ 0.05), ∗∗ (p ≤ 0.01), or ∗∗∗ (p ≤ 0.001), for two-sided p values. (E) Expression of injected xWnt8 variants assessed by western blottingof mid-gastrula-stage embryos. Representative of at least three repeats. (F) Dorsalization phenotypes observed in zebrafish embryos upon ectopic expression of xWnt8WT or xWnt8S187A mRNA. Pictures (top row) show representative embryos at 1 day post fertilization displaying normal (left), moderately dorsalized (“twisted,” center), or highly dorsalized (“bustled,” right) phenotypes. Quantitation of observed phenotypes is shown below, with number of embryos scored across at least two biological replicates listed for each bar. |