Fig. S1
Analysis of Nucleation and Apoptosis in Polyploid Cells (Related to Figure 1)
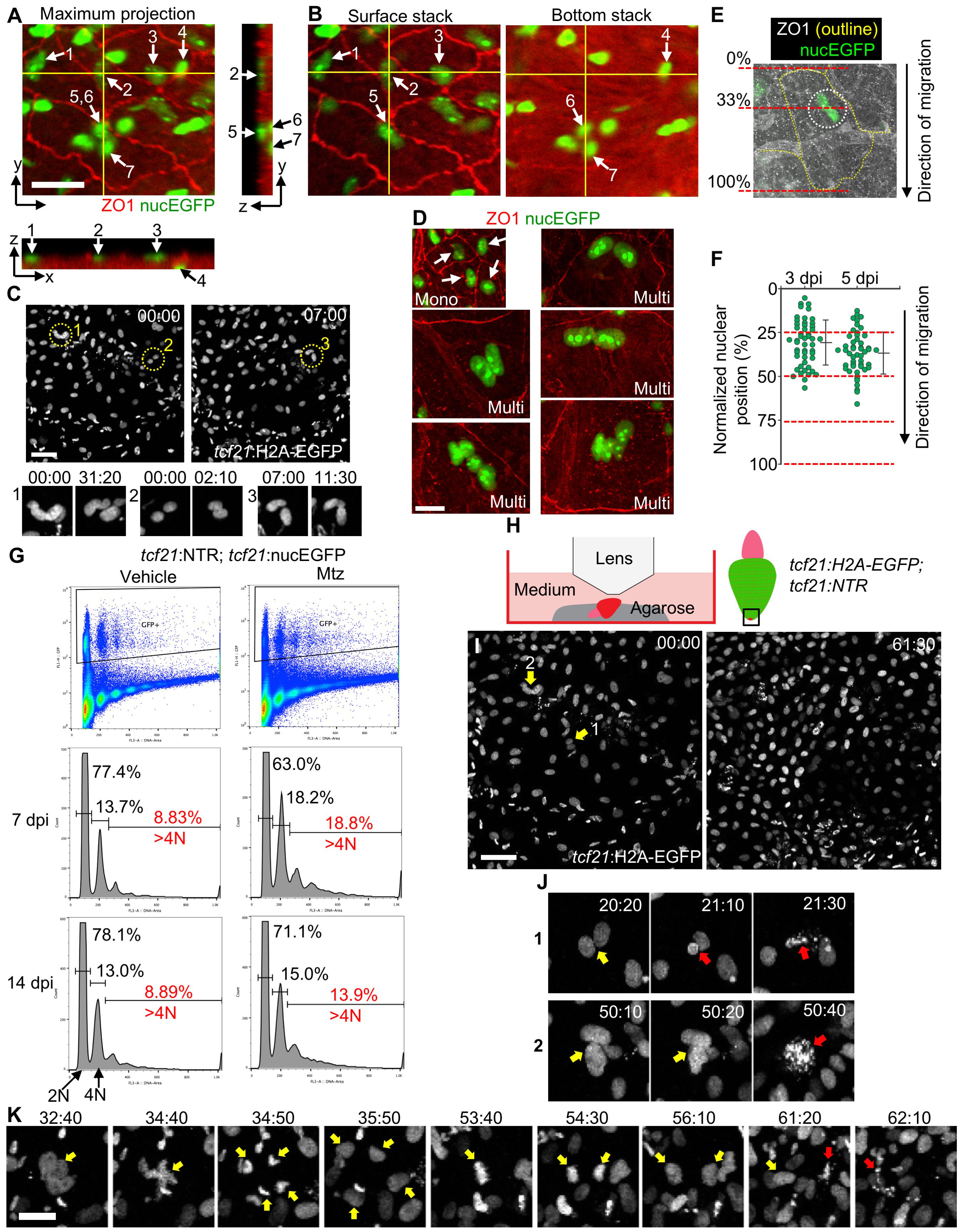
(A) Orthogonal view of a z-stack image. A maximum projection image of the x-y plane is shown with nucEGFP in green and ZO1 staining in red. Yellow lines indicate positions for views of the y-z plane (right) and the x-z plane (bottom), respectively. Arrows and numbers denote nuclei. Scale bar, 20 μm.
(B) Two z-axis stacks of the image shown in (A). The numbers indicate the same nuclei shown in (A). The nuclei 1, 2, 3, and 5 are in the outermost layer, while nuclei 4, 6, and 7 are in the inner layer, embedded within the muscle (red autofluorescence).
(C) Video frames from Movie S1 with tcf21:H2A-EGFP shown in grayscale. N uclei of three cells are circled in low-magnification images (top), and two frames of each are displayed below. Nuclei within individual cells frequently overlap. Scale bar, 50 μm.
(D) Examples of mononucleate cells (top left, arrows) and multinucleate cells. Scale bar, 20 μm.
(E, F) Quantification of nuclear positions within each multinucleate cell at 3 and 5 dpi, respectively. The cell length was measured parallel to the direction of epicardial tissue regeneration. The nuclear position was defined as the center of an area encircling all of the nuclei in each cell and was normalized in percentiles to cell length. n = 50 for each timepoint. Bars indicate mean ± S.D.
(G) FACS analysis of purified epicardial cells. Epicardial cells were purified from uninjured (Vehicle, left) and regenerating samples (Mtz, right) and stained with PI for DNA content. tcf21:NTR; tcf21:nucEGFP ventricles were collected at 7 and 14 dpi. Percentages of 2N, 4N and >4N cells are indicated. Samples from regenerating hearts have 113% (7 dpi) or 56% (14 dpi) more >4N cells (red) than those from uninjured hearts.
(H) Schematic for experiments in (I-K).
(I) Video frames acquired at the ventricular apex at the final stages of epicardial regeneration, covering 61.5 h. tcf21:H2A-EGFP is shown in grayscale. Arrows and numbers denote nuclei shown in (J). Scale bar, 50 μm. See also Movie S1. (J) High-magnification view of nuclear doublets indicated in (I) undergoing nuclear fragmentation (red arrows). Scale bar, 20 μm.
(K) Video frames of an unconventional division of a nuclear doublet undergoing fragmentation. Yellow arrows denote multinucleate cell and daughter nuclei. Red arrows indicate nuclear fragmentation. One daughter nucleus divided again before fragmentation. Scale bar, 20 μm. Timing, hh:mm.
Reprinted from Developmental Cell, 42, Cao, J., Wang, J., Jackman, C.P., Cox, A.H., Trembley, M.A., Balowski, J.J., Cox, B.D., De Simone, A., Dickson, A.L., Di Talia, S., Small, E.M., Kiehart, D.P., Bursac, N., Poss, K.D., Tension Creates an Endoreplication Wavefront that Leads Regeneration of Epicardial Tissue, 600-615.e4, Copyright (2017) with permission from Elsevier. Full text @ Dev. Cell