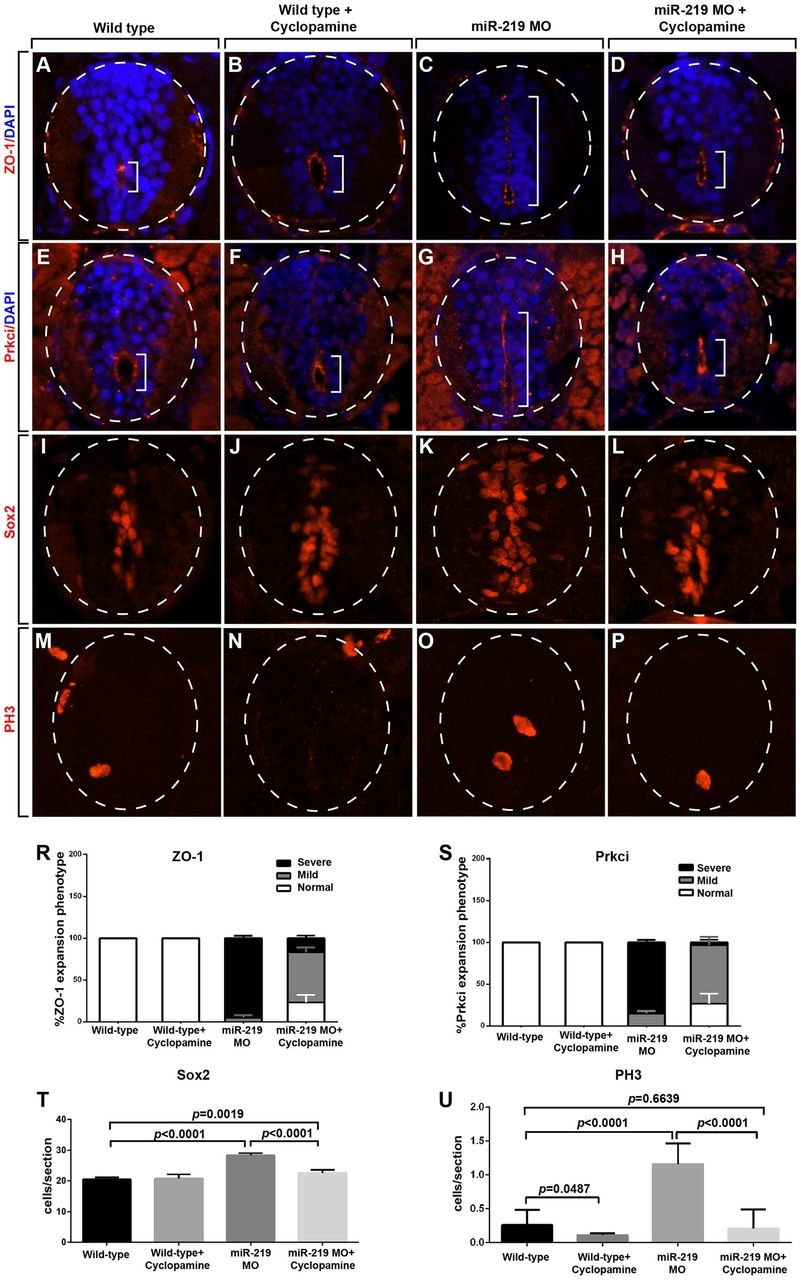
Fig. 2
miR-219-mediated neural progenitor maintenance requires Hh signaling. (A-P) Representative transverse sections through trunk spinal cords of 3dpf embryos with dorsal up. Dashed circles outline spinal cords and brackets indicate central canals/primitive lumens highlighted by ZO-1 and Prkci localization. ZO-1 (A-D) and Prkci (E-H) localization at apical membranes detected by immunohistochemistry. In wild-type (A,E) and cyclopamine-treated wild-type (B,F) embryos, ZO-1 and Prkci are localized to apical membranes surrounding a small, ventrally positioned central canal. miR-219 MO-injected embryos have primitive lumens, decorated by ZO-1 and Prkci, that extend across the dorsoventral length of the spinal cord (C,G). Cyclopamine treatment suppresses the lumenal and apical protein localization phenotype of miR-219 MO-injected embryos (D,H). (I-L) Spinal cord progenitors revealed by Sox2 immunohistochemistry. Cyclopamine treatment suppresses the excess progenitor phenotype of miR-219 MO-injected embryos. (M-P) Dividing spinal cord cells revealed by PH3 immunohistochemistry. Cyclopamine treatment suppresses the excess dividing cell phenotype of miR-219 MO-injected embryos. (R) Graph showing quantification of the ZO-1 phenotype. Embryos classified as normal had ZO-1 expression around the ventrally located central canal. Embryos were classified as severe when ZO-1 localization spanned the entire dorsoventral axis and mild when it spanned an intermediate length. Data represent the mean±s.e.m. (n=15 larvae for each group). P values were calculated by comparing the numbers of larvae with severe and mild phenotypes in miR-219 MO alone and the miR-219 MO+cyclopamine experiments. P<0.0001 for the severe group and P=0.0062 for the mild group, unpaired t-test. (S) Graph showing quantification of the Prkci phenotype. Embryos were scored as in R. Data represent the mean±s.e.m. (n=15 larvae for each group). P<0.001 for the severe group and P=0.0033 for the mild group, unpaired t-test. (T) Graph showing the number of Sox2+ progenitors. Data represent the mean±s.e.m. (n=15 embryos per group). Significance calculated using an unpaired t-test. (U) Graph showing the number of PH3+ cells in wild-type control, wild-type+cyclopamine, miR-219 MO and miR-219 MO+cyclopamine larvae. Data represent the mean±s.e.m. (n=15 sections obtained from 5 larvae per group, with three replicates). Significance calculated using an unpaired t-test.