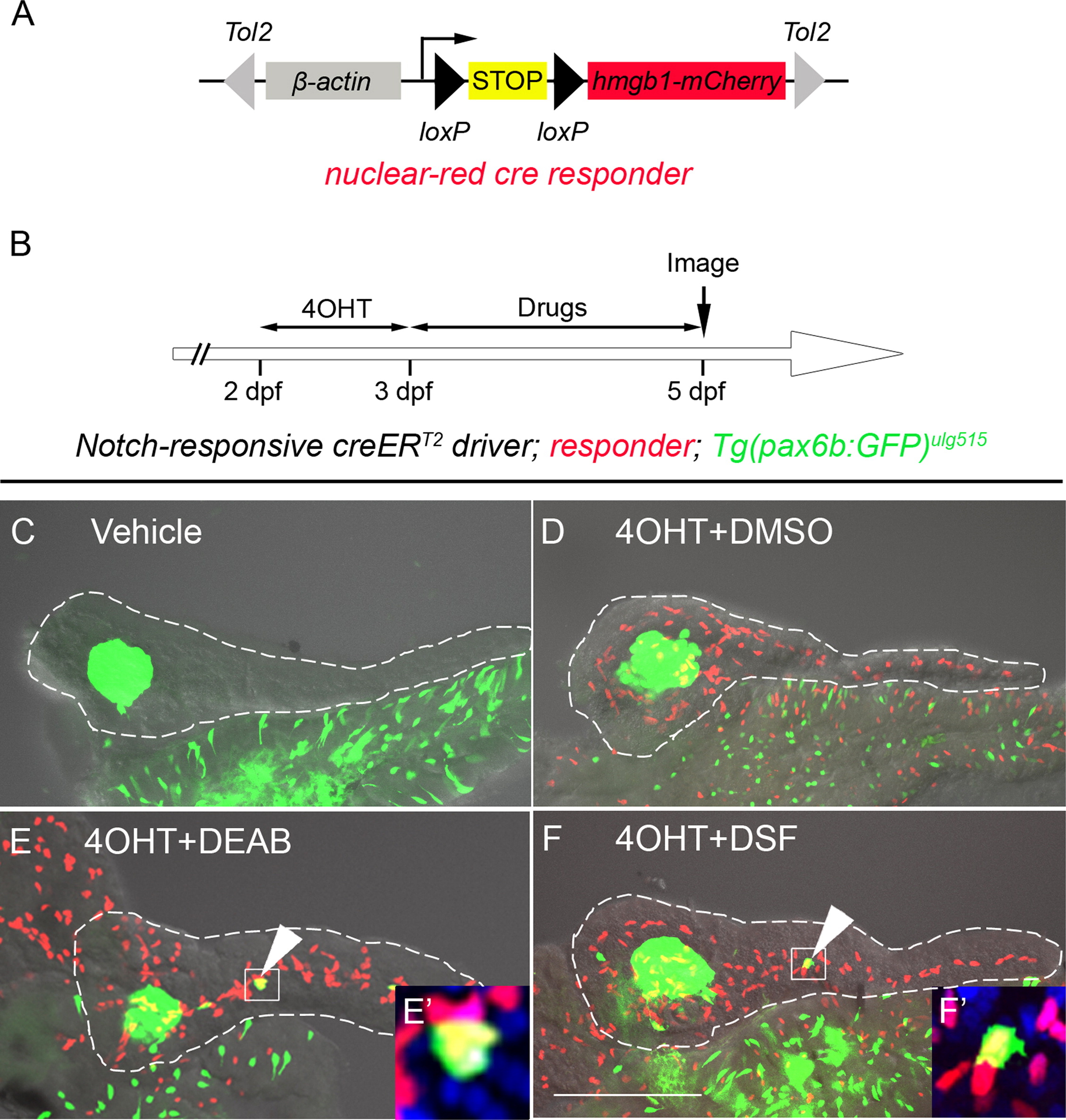
Fig. 3
Inhibition of RA signaling induces PNCs to differentiate into endocrine cells. (A) Schematic of the nuclear red cre responder, Tg(β-actin:loxP-stop-loxP-hmgb1-mCherry)jh15. The β-actin promoter/enhancer drives ubiquitous expression. Active Cre induces recombination between the two loxP sites (black triangles) removing the STOP signal (yellow box) and allowing production of Hmgb1-mCherry, a nuclear-red fluorescent protein. The transgene is flanked by Tol2 arms (gray triangles). (B) Experimental timeline of 4OHT-dependent fate mapping. 4OHT was applied 2–3 dpf. Larvae were then exposed to drugs from 3–5 dpf. (C–F) Confocal z-stack projections of dissected pancreata from larvae transgenic for the Notch-responsive creERT2driver, Tg(Tp1glob:creERT2)jh12, the nuclear red cre responder and the endocrine marker, Tg(pax6b:eGFP)ulg515. These triple transgenic larvae were treated with vehicle (C), 4OHT+DMSO (D), 4OHT+DEAB (E), and 4OHT+DSF (F). The presence of lineage-traced cells (red nuclei) is dependent on 4OHT treatment (D–F). 2° islet cells can be detected in the tail region of larval pancreata after treatment with either DEAB (E) or DSF (F). (E′ and F′) The precocious 2° islet cells are positive for both GFP and mCherry, indicating their PNC origin. Nuclei are counterstained with DAPI (blue). White dashes outline pancreata. Scale Bar, 100 μm.
Reprinted from Developmental Biology, 394(1), Huang, W., Wang, G., Delaspre, F., Vitery, M.D., Beer, R.L., Parsons, M.J., Retinoic acid plays an evolutionarily conserved and biphasic role in pancreas development, 83-93, Copyright (2014) with permission from Elsevier. Full text @ Dev. Biol.