- Title
-
Virulomic Analysis of Multidrug-Resistant Klebsiella pneumoniae Isolates and Experimental Virulence Model Using Danio rerio (Zebrafish)
- Authors
- Duarte, E.L.T., Rizek, C.F., Espinoza, E.S., Marchi, A.P., Noguera, S.V., Côrtes, M.F., Fernandes, B.H.V., Guimarães, T., de Maio Carrilho, C.M.D., Neto, L.V.P., Trindade, P.A., Costa, S.F.
- Source
- Full text @ Antibiotics (Basel)
|
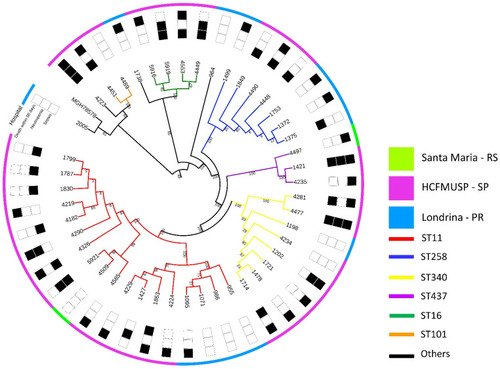
Phylogenetic tree of all 46 K. pneumoniae isolates in this study, based on single-nucleotide polymorphisms (SNPs) of core genome. Colors on the outermost circle represent place of origin, while branch colors represent sequence types. Each number represents one strain, with each isolated from a different patient. |
|
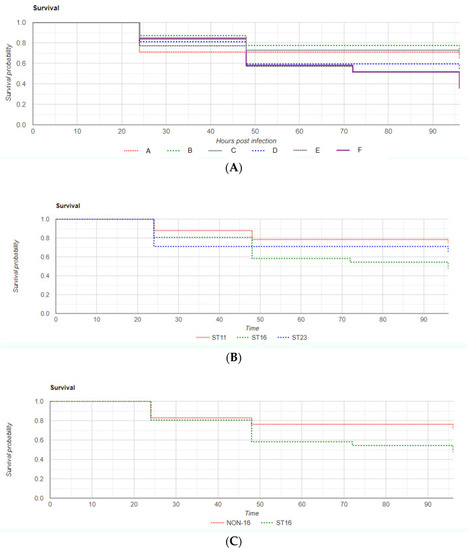
Overall zebrafish Kaplan–Meier survival curve (A) according to K. pneumoniae strain of infection; (B) Kaplan–Meier curves when grouping survival of embryos by ST of infection; (C) Kaplan–Meier survival curves comparing zebrafish infection by K. pneumoniae ST16 strains with non-ST16 strains. Time is expressed in hours post-infection (hpi). PHENOTYPE:
|