- Title
-
Tryptophan Hydroxylase 2 Deficiency Modifies the Effects of Fluoxetine and Pargyline on the Behavior, 5-HT- and BDNF-Systems in the Brain of Zebrafish (Danio rerio)
- Authors
- Evsiukova, V.S., Bazovkina, D., Bazhenova, E., Kulikova, E.A., Kulikov, A.V.
- Source
- Full text @ Int. J. Mol. Sci.
|
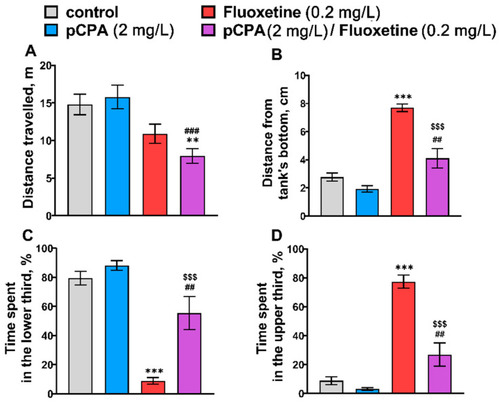
(A) Distance traveled (m), (B) mean distance from the tank’s bottom (cm), time (%) (C) spent in the lower and (D) the upper thirds in the novel tank diving test in control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/L), fluoxetine (0.2 mg/L) or combination of pCPA (2 mg/L) and fluoxetine (0.2 mg/L), correspondently. The number of animals in each group was 10. ** p < 0.01, *** p < 0.001 vs. control group; ## p < 0.01, ### p < 0.001 vs. pCPA treated group; $$$ p < 0.001 vs. fluoxetine treated group. |
|
Concentration of (A) 5-HT (ng/mg of protein), (B) 5-HIAA (ng/mg of protein) and (C) 5-HIAA/5-HT ratio in the brain of control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/L), fluoxetine (0.2 mg/L) or combination of pCPA (2 mg/L) and fluoxetine (0.2 mg/L), correspondently. The number of animals in each group was 10. * p < 0.05, ** p < 0.01, *** p < 0.001 vs. control group; ### p < 0.001 vs. pCPA treated group; $$$ p < 0.001 vs. fluoxetine treated group. |
|
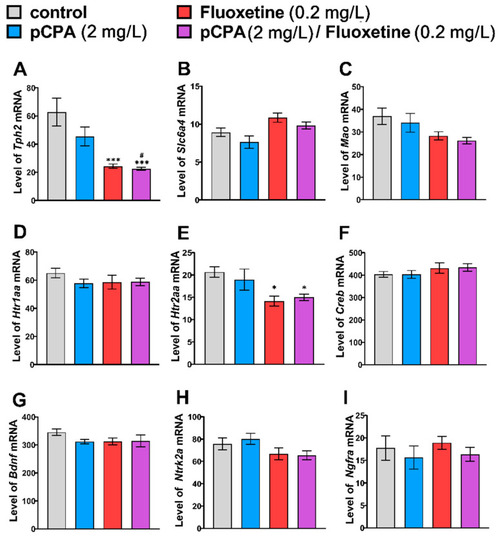
Levels of mRNA of (A) Tph2, (B) Slc6a4b, (C) Mao, (D) Htr1aa, (E) Htr2aa, (F) Creb, (G) Bdnf, (H) Ntrk2a and (I) Ngfra genes in the brain of control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/L), fluoxetine (0.2 mg/L) or combination of pCPA (2 mg/L) and fluoxetine (0.2 mg/L), correspondently. The number of animals in each group was 10. * p < 0.05, *** p < 0.001 vs. control group; # p < 0.05 vs. pCPA treated group. |
|
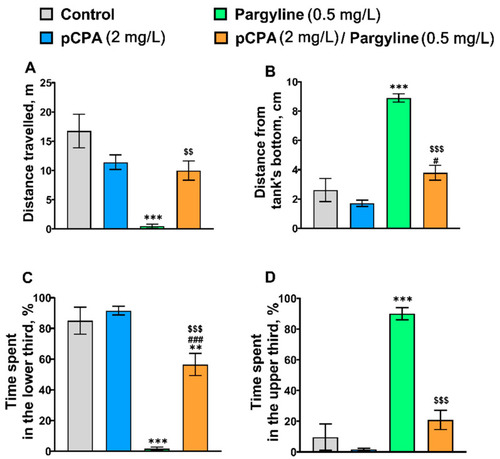
(A) Distance traveled (m), (B) mean distance from the tank’s bottom (cm), (C) time (%) spent in the lower and (D) the upper thirds in the novel tank diving test in control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/L), pargyline (0.5 mg/L) or combination of pCPA (2 mg/L) and pargyline (0.5 mg/L), correspondently. The number of animals in each group was 10. ** p < 0.01; *** p < 0.001 vs. control group; # p < 0.05, ### p < 0.001 vs. pCPA treated group; $$ p < 0.01, $$$ p < 0.001 vs. pargyline treated group. |
|
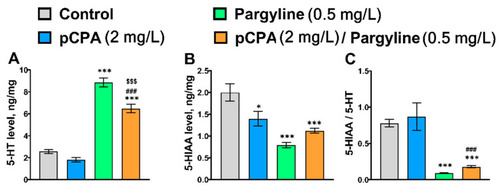
Concentration of (A) 5-HT (ng/mg of protein), (B) 5-HIAA (ng/mg of protein) and (C) 5-HIAA/5-HT ratio in the whole brain of control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/L), pargyline (0.5 mg/L) or combination of pCPA (2 mg/L) and pargyline (0.5 mg/L), correspondently. The number of animals in each group was 10. * p < 0.05, *** p < 0.001 vs. control group; ### p < 0.001 vs. pCPA treated group; $$$ p < 0.001 vs. pargyline treated group. |
|
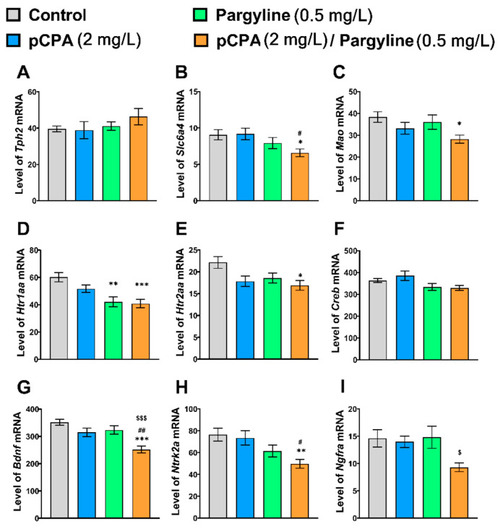
Levels of mRNA of (A) Tph2, (B) Slc6a4b, (C) Mao, (D) Htr1aa, (E) Htr2aa, (F) Creb, (G) Bdnf, (H) Ntrk2a and (I) Ngfra genes in the whole brain of control (water) zebrafish and those exposed for 72 h to pCPA (2 mg/l), pargyline (0.5 mg/L) or combination of pCPA (2 mg/L) and pargyline (0.5 mg/L), correspondently. The number of animals in each group was 10. * p < 0.05, ** p < 0.01, *** p < 0.001 vs. control group; # p < 0.05, ## p < 0.01 vs. pCPA treated group; $ p < 0.05, $$$ p < 0.001 vs. pargyline treated group. |
|
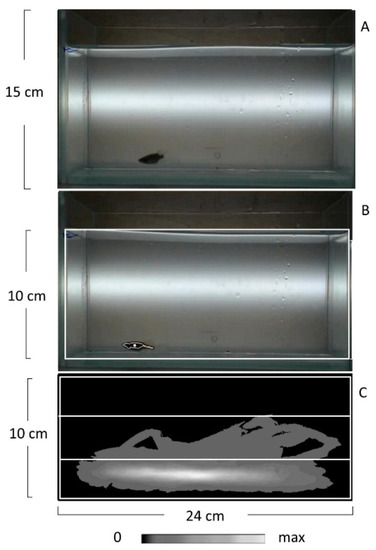
Tracking zebrafish in the novel tank diving test by EthoStudio software. (A) Photo of the tank for testing (24 cm in length, 15 cm in depth, and 7 cm in width) filled with water and with a zebrafish inside. (B) The white rectangle marks the water surface selected as the tracking arena, which is 24 cm in length and 10 cm in depth. EthoStudio software finds the contour and the center of a zebrafish and marks them with a white line and a point, respectively. (C) EthoStudio software calculates the density map of pixel distribution associated with zebrafish on the arena. Pixel density is coded gray according to the scale placed below the map. The arena is divided into three equal thirds. The map shows that most of the pixels associated with a zebrafish are in the lower third. |