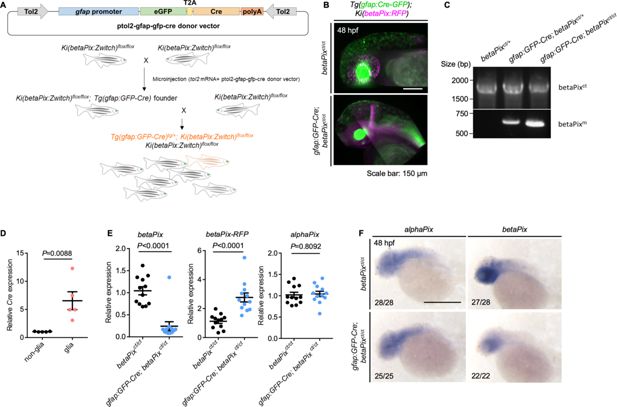
Fig. 3 - Supplemental 1 Generating glial-specific betaPix knockout zebrafish. (A) Schematic diagram illustrates establishment of glial-specific betaPix knockout zebrafish. (B) 3D reconstruction of the heads at 48 hpf with lateral view, anterior to left. Glial-specific Cre expression (green) was only shown in gfap:GFP-Cre; betaPixct/ct mutant embryos, which overlaps with betaPix:RFP expression (magenta). (C) Genomic PCR analysis of the betaPixct- or betaPixm-unique sequences in betaPixct/ct siblings, gfap:GFP-Cre; betaPixct/+ heterozygous mutants, and gfap:GFP-Cre; betaPixct/ct homozygous mutants. (D) qRT-PCR analysis revealing Cre expression in gfap:GFP-positive glial population and gfap:GFP-negative non-glial population that were sorted from gfap:GFP-Cre; betaPixct/ct embryos. Each dot represents one embryo. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (E) qRT-PCR analysis showing betaPix, betaPix-RFP, and alphaPix expression in betaPixct/ct control siblings and gfap:GFP-Cre; betaPixct/ct mutant embryos at 48 hpf. Each dot represents one embryo. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (F) Whole-mount RNA in situ hybridization revealing that alphaPix was comparable but betaPix decreased in gfap:GFP-Cre; betaPixct/ct mutants compared with betaPixct/ct siblings at 48 hpf. Lateral view with anterior to the left. Individual scale bars are indicated in the figure.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Elife