Figure 2
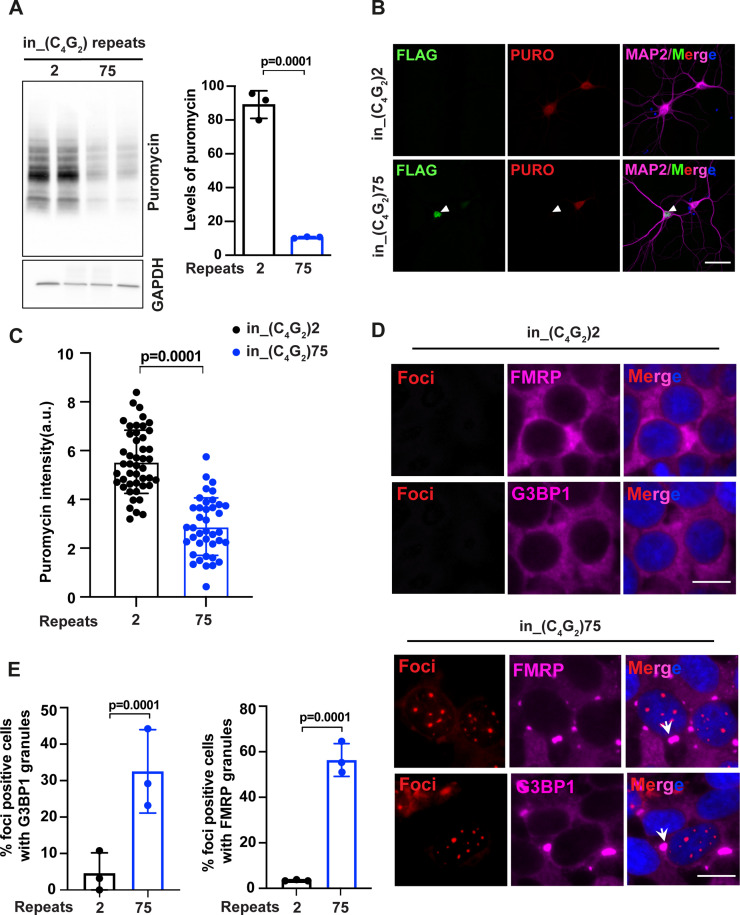
(A) Immunoblotting of puromycin in HEK293T cells expressing in_(C4G2)75 or 2 repeats. Cells were incubated with puromycin for 30 min before harvesting. The level of puromycin was normalized to GAPDH. Error bars represent SD (n = 3 independent experiments). Statistical analyses were performed using Student’s t-test (B) Representative images and (C) quantification of primary neurons expressing either (C4G2)75 or 2 repeats stained with anti-puromycin (red), anti-FLAG (green), DAPI (blue), and MAP2 (magenta). The puromycin intensity was quantified using ImageJ. Error bars represent SD (n = 40–50 neurons/group; similar results were obtained from two independent experiments). Statistical analyses were performed using Student’s t-test. (D) Representative images of G3BP1 and FMRP staining in HEK293T cells expressing in_(C4G2)75 and in_(C4G2)2 repeats. (E) Quantification of antisense foci-positive cells with G3BP1 and FMRP granules. Error bars represent SD (n = 150 cells/condition and three independent experiments). Statistical analyses were performed using Student’s t-test. Scale bars, 10 µm (neurons), 20 µm (HEK293T).
C9ORF72 antisense C4G2 expanded repeats inhibit global protein synthesis and induce stress granule assembly.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Elife