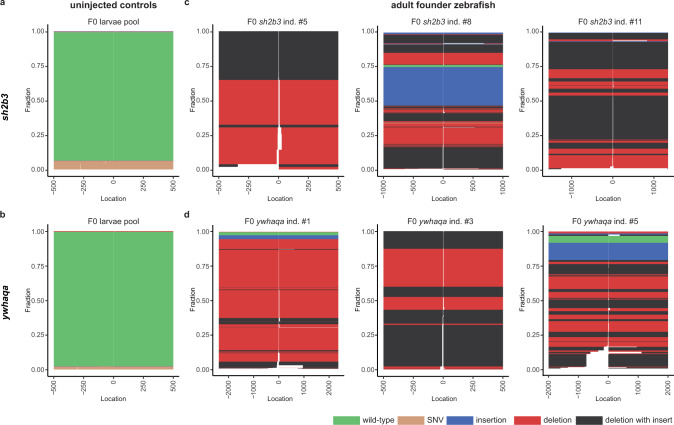
Fig. 4 Schematic view of sequencing reads and editing events at the on-target sites for sh2b3 and ywhaqa, produced by the SIQ software. In each plot, the y-axis displays the fraction of different editing events, and the x-axis their genomic positions. The zero coordinate indicates the Cas9-cleavage site for sh2b3 and ywhaqa. Control samples from uninjected zebrafish displaying the sh2b3 (a) and ywhaqa (b) targeted sites. Only wild-type alleles and sequences containing SNVs are detected in these controls. SIQ results from three individual F0 founder fish edited for sh2b3 (c) or ywhaqa (d). In the CRISPR-Cas9 edited F0 individuals, a large number of distinct insertions, deletions, as well as combinations of insertions and deletions are reported. This indicates a high degree of somatic mosaicism of CRISPR-Cas9 editing events in the F0 individuals. Source data are provided as a Source Data file.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Nat. Commun.