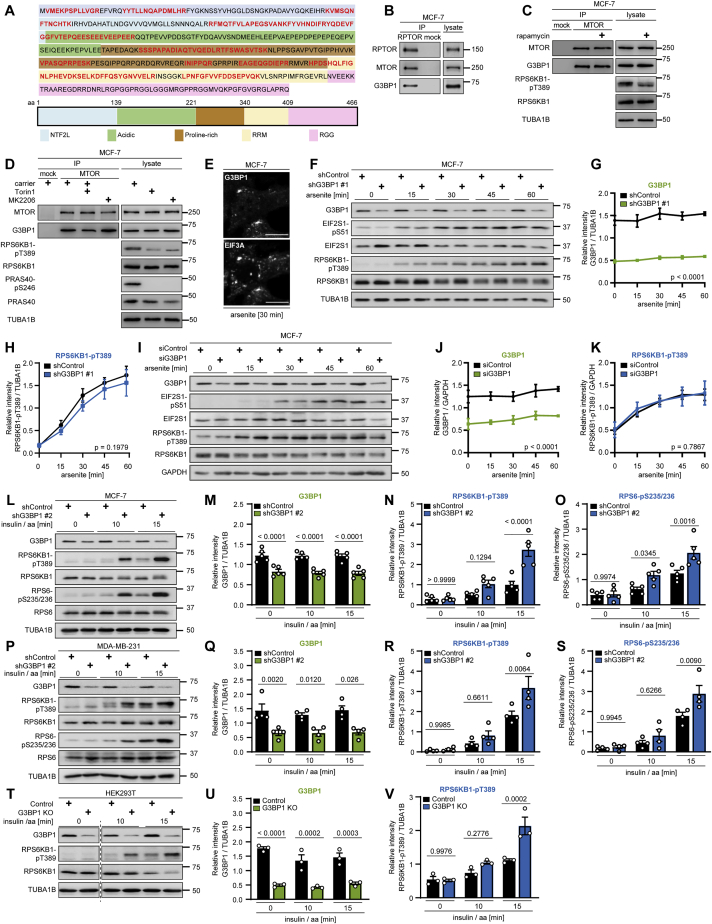
Figure S1
G3BP1 does not alter mTORC1 activity upon arsenite stress but upon stimulation with insulin and nutrients, related to
(A) Amino acid sequence of G3BP1. Protein domains indicated according to
(B) IP against RPTOR (RPTOR#1 or #2) or mock (rat IgG). n = 3.
(C) IP against MTOR or mock (rat IgG) from rapamycin-treated cells. n = 3.
(D) IP against MTOR or mock (rat IgG) from Torin1 or MK2206treated cells. n = 3.
(E) IF analysis of G3BP1 and EIF3A in arsenite exposed cells. Scale bar, 10 μm. n = 3.
(F) Time course analysis of shG3BP1 #1 cells exposed to arsenite for up to 60 min. n = 3.
(G) Quantitation of G3BP1 in (F). Mean ± SEM.
(H) Quantitation of RPS6KB1-pT389 in (F). Data shown as in (G).
(I) Time course analysis of siG3BP1 cells exposed to arsenite for up to 60 min. n = 3.
(J) Quantitation of G3BP1 in (I). Mean ± SEM.
(K) Quantitation of RPS6KB1-pST389 in (I). Data shown as in (J).
(L) Insulin and amino acid (insulin/aa)-stimulated G3BP1 knockdown cells harboring a second shRNA sequence (shG3BP1 #2) targeting another exon than shG3BP1 #1 (
(M) Quantitation of G3BP1 in (L). Shown are data points and mean ± SEM.
(N) Quantitation of RPS6KB1-pT389 in (L). Data shown as in (M).
(O) Quantitation of RPS6-pS235/236 in (L). Data shown as in (M).
(P) Insulin/aa-stimulated shG3BP1 #2. n = 4.
(Q) Quantitation of G3BP1 in (P). Shown are data points and mean ± SEM.
(R) Quantitation of RPS6KB1-pT389 in (P). Data shown as in (Q).
(S) Quantitation of RPS6-pS235/236 in (P). Data shown as in (Q).
(T) Insulin/aa-stimulated G3BP1 KO cells generated with a second independent guide RNA against G3BP1 (sgRNA # 2,
(U) Quantitation of G3BP1 in (T). Shown are data points and mean ± SEM.
(V) Quantitation of RPS6KB1-pT389 in (T). Data shown as in (U).
Reprinted from Cell, 184(3), Prentzell, M.T., Rehbein, U., Cadena Sandoval, M., De Meulemeester, A.S., Baumeister, R., Brohée, L., Berdel, B., Bockwoldt, M., Carroll, B., Chowdhury, S.R., von Deimling, A., Demetriades, C., Figlia, G., Genomics England Research Consortium, de Araujo, M.E.G., Heberle, A.M., Heiland, I., Holzwarth, B., Huber, L.A., Jaworski, J., Kedra, M., Kern, K., Kopach, A., Korolchuk, V.I., van 't Land-Kuper, I., Macias, M., Nellist, M., Palm, W., Pusch, S., Ramos Pittol, J.M., Reil, M., Reintjes, A., Reuter, F., Sampson, J.R., Scheldeman, C., Siekierska, A., Stefan, E., Teleman, A.A., Thomas, L.E., Torres-Quesada, O., Trump, S., West, H.D., de Witte, P., Woltering, S., Yordanov, T.E., Zmorzynska, J., Opitz, C.A., Thedieck, K., G3BPs tether the TSC complex to lysosomes and suppress mTORC1 signaling, 655-674.e27, Copyright (2021) with permission from Elsevier. Full text @ Cell