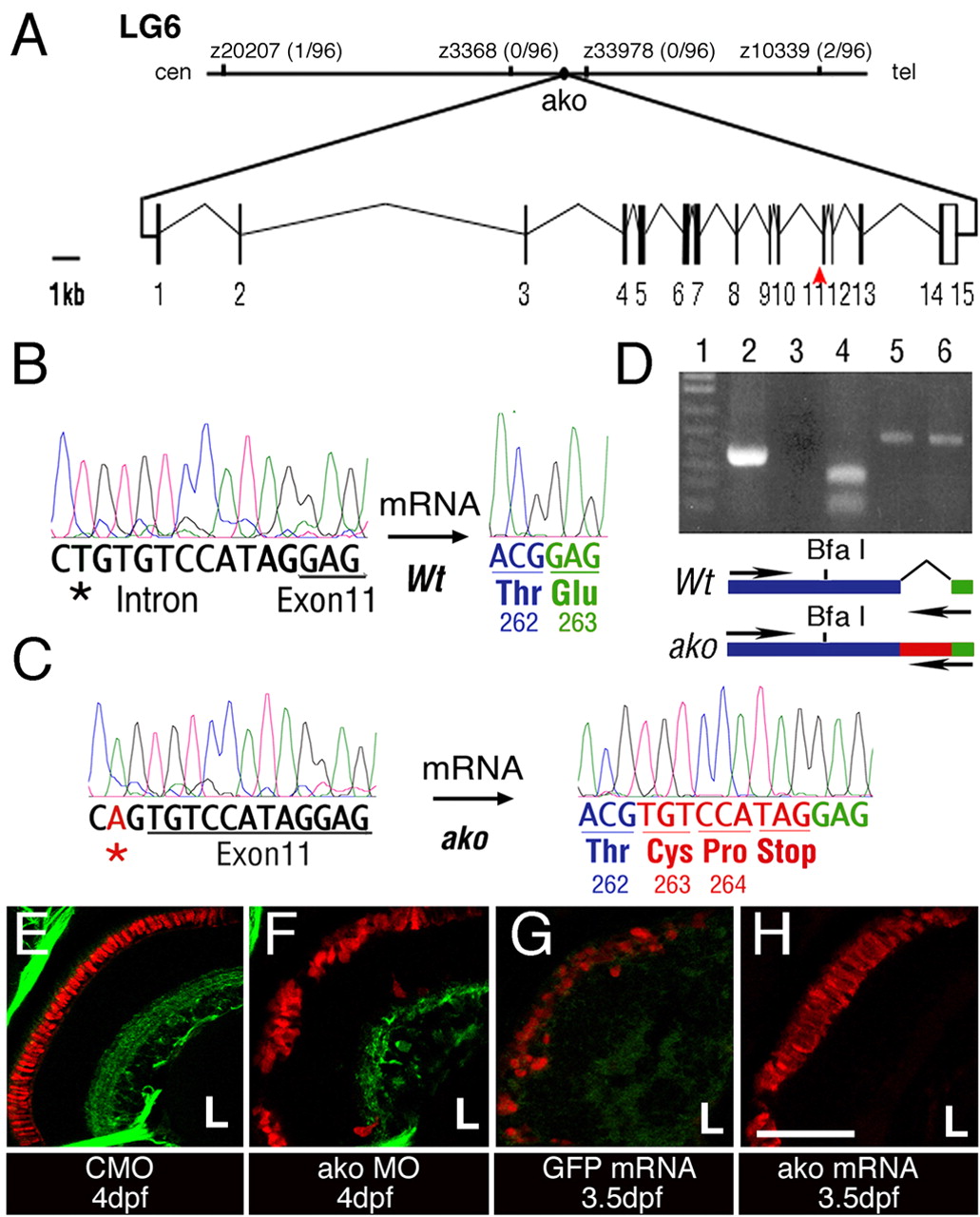
Fig. 3 The ale oko locus encodes a zebrafish dynactin 2 homolog. (A) Map of the ako genomic region and exon/intron structure of the ako transcript. The position of mutant sequence in the akojj50 allele is indicated (red arrowhead). (B,C) Comparison of sequence trace data from wild-type (B) and mutant (C) animals. The akojj50 mutation activates a cryptic splice site in the genomic sequence. This, in turn, introduces a 9-bp insertion (red) that encodes a premature stop codon. (D) RT-PCR amplification of the dynactin 2 transcript using a primer that recognizes the mutant insertion sequence. Lane 1, DNA ladder; lane 2, amplification of the mutant transcript produces a single robust band; lane 3, amplification of the wild-type transcript does not produce a detectable product; lane 4, BfaI digestion confirms the identity of the amplification product from the mutant; lanes 5 and 6, control PCR amplification of actin from wild-type (5) and mutant (6) animals. The positions of amplification primers in the wild type and mutant are indicated on the diagram below. (E-H) Phenocopy and rescue of photoreceptor defects in ako mutant animals. Transverse cryosections through embryonic retinae were stained with the Zpr-1 antibody to visualize double cones (red). Green signal in E,F originates from the Tg(brn3c:mGFP) transgene. The injection of ako (H), but not GFP (G), mRNA rescues photoreceptor morphology in ako mutants. L, lens. Scale bar: 40 μm.
Image
Figure Caption
Figure Data
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Development