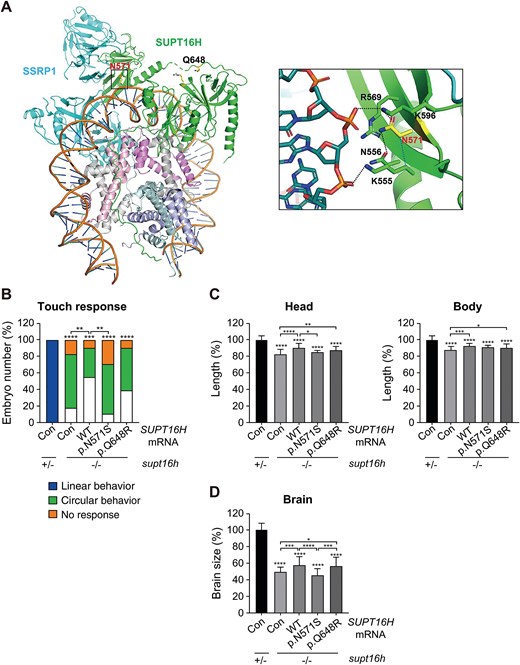
Fig. 3 Structural and functional analysis of SUPT16H variants. (A) Positions of the two SUPT16H variants within the structure of the FACT–nucleosome complex (PDB ID: 6UPK). The side chains of N571 (labeled in red) and Q648 (labeled in blue) are shown as yellow sticks. SUPT16 and SSRP1 are depicted in green and cyan, respectively, along with DNA bound to the (H3–H4)2 tetramer and the H2A–H2B dimer. Right: Close-up view of the polar interaction network surrounding N571, with polar contacts indicated by dotted lines (see also Figs. S2A–C). (B) Touch-evoked swimming patterns in embryos injected with human SUPT16H mRNA (WT, c.1712A > G [p.N571S], or c.1943A > G [p.Q648R]) assessed at 2 dpf. The control group (con) was injected with TagRFP mRNA. Data are presented as mean values; n ≥ 45 embryos per group. (C) Bright-field images of embryos injected with SUPT16H mRNA. Head size and body length were measured at 2 dpf (see also Fig. S2D). Data are presented as mean ± SD; n ≥ 10 embryos per group. (D) Brain morphology of embryos injected with SUPT16H mRNA was assessed by anti-acetylated tubulin immunohistochemistry to visualize neuronal structures (see also Fig. S2E). Brain size was quantified at 2.5 dpf. Data are presented as mean ± SD; n ≥ 25 embryos per group. Head size, body length, and brain size were measured using ImageJ software. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparison test; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Abbreviations: ANOVA, analysis of variance; Con, control (TagRFP mRNA-injected); CTD, C-terminal domain; DD, dimerization domain; dpf, days post-fertilization; FACT, FAcilitates Chromatin Transcription; MD, middle domain; NTD, N-terminal domain; PDB, Protein Data Bank; SD, standard deviation; SUPT16H, suppressor of Ty 16 protein; WT, wild type.
Image
Figure Caption
Figure Data
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Hum. Mol. Genet.