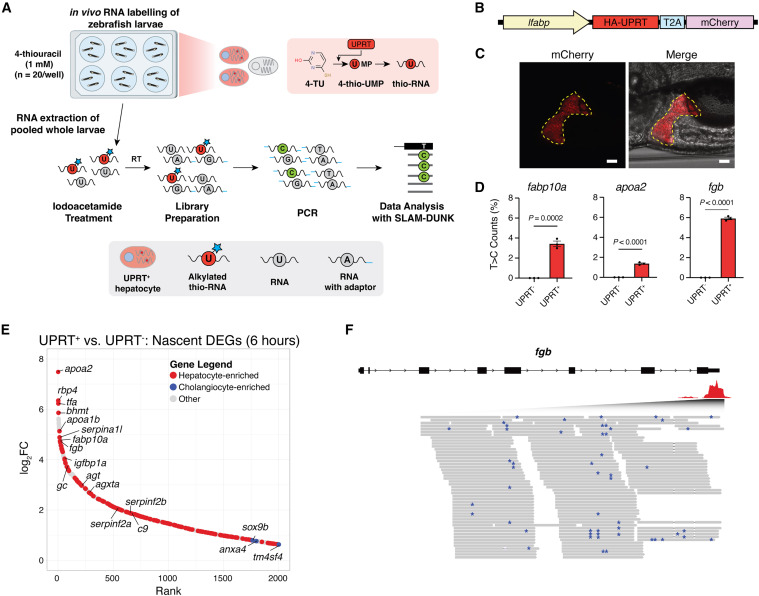
Fig. 1 Development of a transgenic zebrafish model for the identification of hepatocyte-specific nascent gene expression (A) Schematic of larval SLAM-ITseq workflow. (B) Tg(lfabp:UPRT-T2A-mCherry) construct. (C) Fluorescent validation of hepatocyte-specific mCherry expression in 5 dpf zebrafish larvae. Yellow dashed line borders the mCherry+ liver. Scale bars, 50 μm. (D) Percentage of T>C counts for hepatocyte-enriched nascent transcripts (fabp10a, apoa2, and fgb) in UPRT+ and UPRT− zebrafish labeled with 4TU for 6 h. p values were calculated using an unpaired two-tailed Student’s t test comparing each sample to the respective UPRT− control. (E) Hockey plot of nascent differentially expressed genes (DEGs) ranked by log-fold change comparing UPRT+ and UPRT− zebrafish labeled with 4TU for 6 h. Red genes indicate hepatocyte-enriched transcripts and blue genes indicate cholangiocyte-enriched transcripts obtained from the Human Protein Atlas (proteinatlas.org ). (F) Representative screenshot of the Integrative Genomics Viewer (IGV) genome browser within the 3′ UTR locus of fgb derived from 3′ RNA SLAM-ITseq. Blue asterisks indicate T>C conversions. See also Figure S1.
Reprinted from Developmental Cell, 59(7), Tan, V.W.T., Salmi, T.M., Karamalakis, A.P., Gillespie, A., Ong, A.J.S., Balic, J.J., Chan, Y.C., Bladen, C.E., Brown, K.K., Dawson, M.A., Cox, A.G., SLAM-ITseq identifies that Nrf2 induces liver regeneration through the pentose phosphate pathway, 898-910.e6, Copyright (2024) with permission from Elsevier. Full text @ Dev. Cell