Figure 7
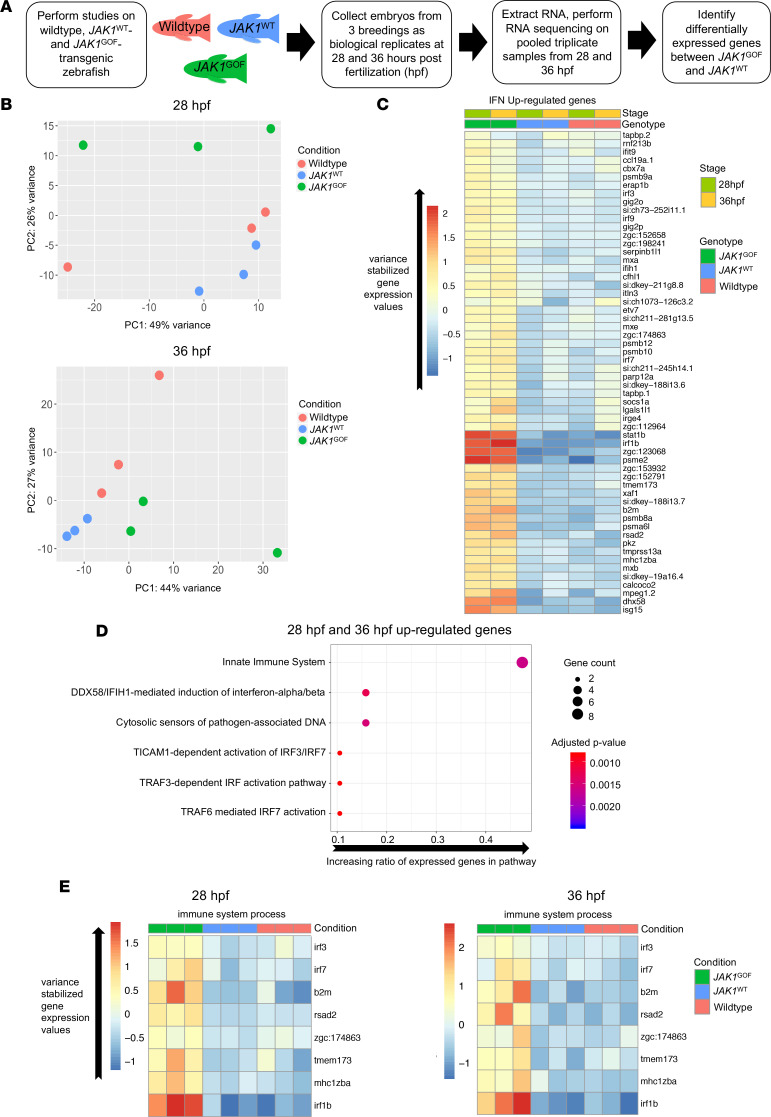
(A) Description of steps involved in the RNA-Seq experiment aimed at identifying genes specifically induced by JAK1-A634D (JAK1GOF) in zebrafish embryos. After performing standard sample collection and RNA-Seq steps, the main comparison at the analysis stage was between JAK1WT and JAK1GOF samples. (B) PCA of 28 and 36 hpf data sets consisting of WT, JAK1WT, and JAK1GOF samples. The plots shown contain the first 2 components (PC1, PC2) and identify clear differences between JAK1GOF samples and JAK1WT or uninjected samples at 28 hpf. (C) Heatmap of average variance-stabilized gene expression values for IFN-stimulated genes that are present among the genes upregulated by JAK1GOF in at least 1 stage analyzed (28 and 36 hpf) (58 genes) in all analyzed groups of samples. (D) Reactome Pathway analysis of genes upregulated by JAK1GOF shows strong enrichment of immunity-related and DNA damage pathways. Gene ratio is the fraction of the genes in a pathway that were present in the input gene list. Gene count and P values associated with a pathway are indicated by the point size and its color. (E) Heatmaps of average variance-stabilized gene expression values for genes associated with the “immune system process” GO term in all analyzed groups of samples.