Figure 4
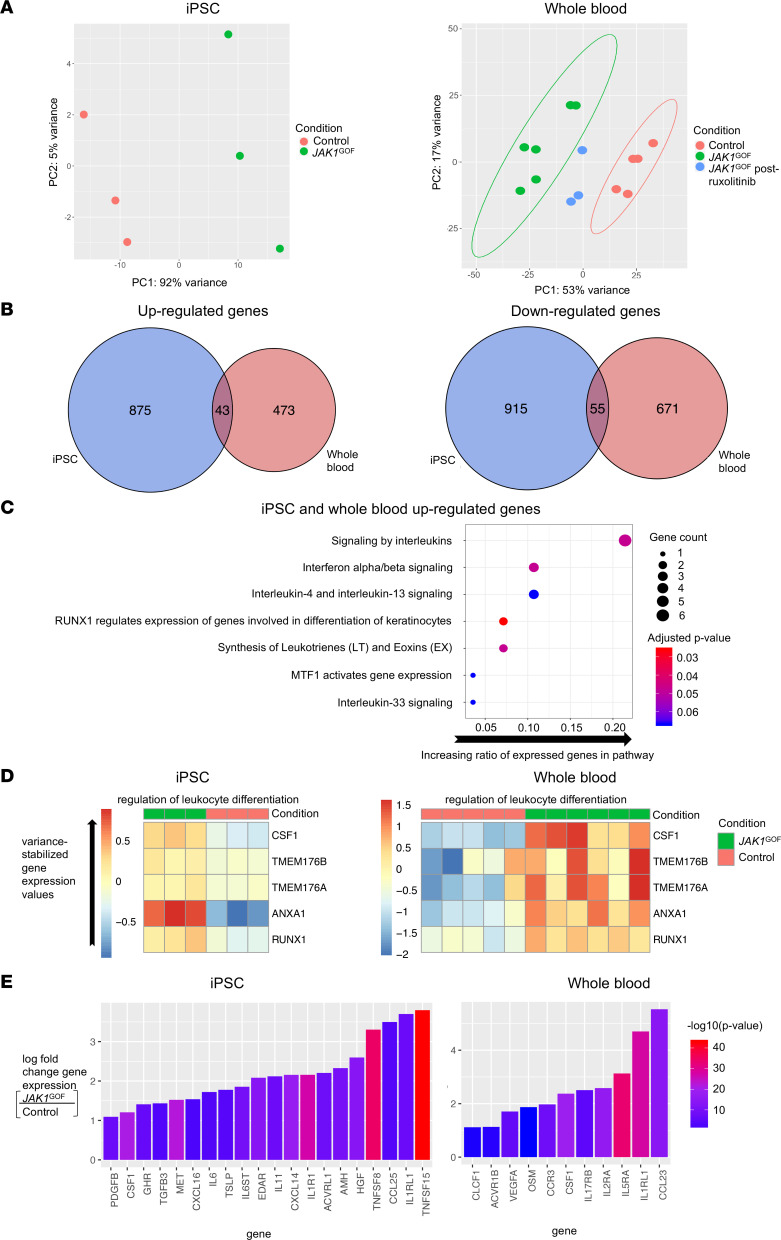
(A) PCA of iPSC and whole blood showing clustering of JAK1GOF compared with controls, with JAK1GOF samples from patients following treatment with ruxolitinib shifting closer to controls. Ellipses demonstrating independent clustering are shown for groups containing more than 3 samples. (B) Analysis of differentially expressed genes in human iPSC and whole blood carrying the JAK1GOF demonstrated 43 upregulated and 55 downregulated genes in both iPSC and whole blood compared with controls. (C) Curated list of significantly enriched pathways performed by Reactome Pathway analysis in the 43 upregulated genes found in both human JAK1GOF iPSC and whole blood samples. The y axis indicates Reactome Pathway, and the x axis corresponds to ratio of upregulated genes to the number of genes in each pathway, and the size of each dot corresponds to number of genes identified in samples. Color corresponds to the adjusted P value, with red being more significant. (D) Heatmap of variance-stabilized gene expression values from ontology term groups that were enriched in the shared upregulated genes in human JAK1GOF iPSC and whole blood samples. (E) Waterfall plots of cytokine and cytokine receptors upregulated in human JAK1GOF iPSC and whole blood samples. The y axis corresponds to the log fold change of JAK1GOF compared with controls, and the x axis corresponds to gene name; colors indicate –log(P value), with red being more significant. Whole blood RNA-Seq data obtained from 5 healthy controls, 3 JAK1GOF patients (2 samples each from II-2, III-1, III-2) obtained at different time points before treatment, and 2 JAK1GOF patients (III-1, III-2) after treatment (1 sample from III-1 and 2 samples from III-2 obtained at different time points). iPSC RNA-Seq data were obtained from iPSC-derived EBs on day 7 of differentiation (3 samples each from JAK1GOF and Ctrl, with 50 EBs collected per sample).