Fig 9
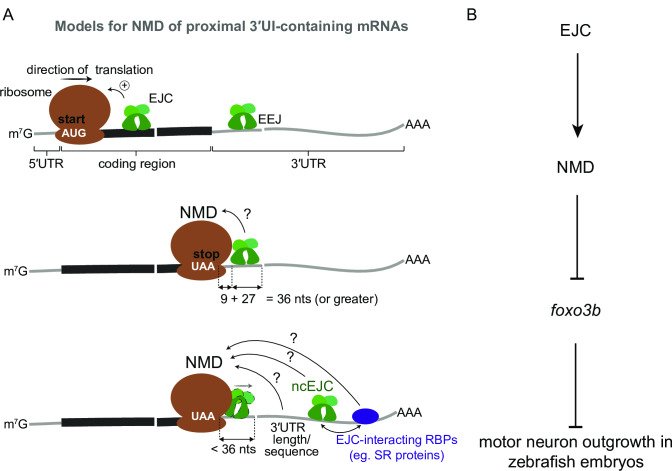
A. Top: EJCs can function to enhance NMD via translation stimulation. Ribosome (brown), direction of translation (straight arrow), start codon (‘AUG’ in white), EJC (green), exon-exon junction (EEJ), mRNA coding region (thick black line) and mRNA 5′ and 3′UTRs (thin gray lines) are labeled. Middle: a model for downstream EJC function in NMD of transcripts where the distance between stop codon (‘UAA’ in white) and downstream exon-exon junction is 36 nts or greater to accommodate both the EJC and the ribosome. Bottom: a model for NMD of transcripts where the distance between stop codon and downstream exon-exon junction is less than 36 nts. Displacement of stop codon-proximal EJC by the ribosome is shown. In this case other downstream factors such as a non-canonical EJC (ncEJC), EJC-interacting RBPs (e.g. SR proteins), or 3′UTR length and/or specific sequences may be responsible for NMD. B. A schematic depicting a model for EJC- and NMD-dependent regulation of