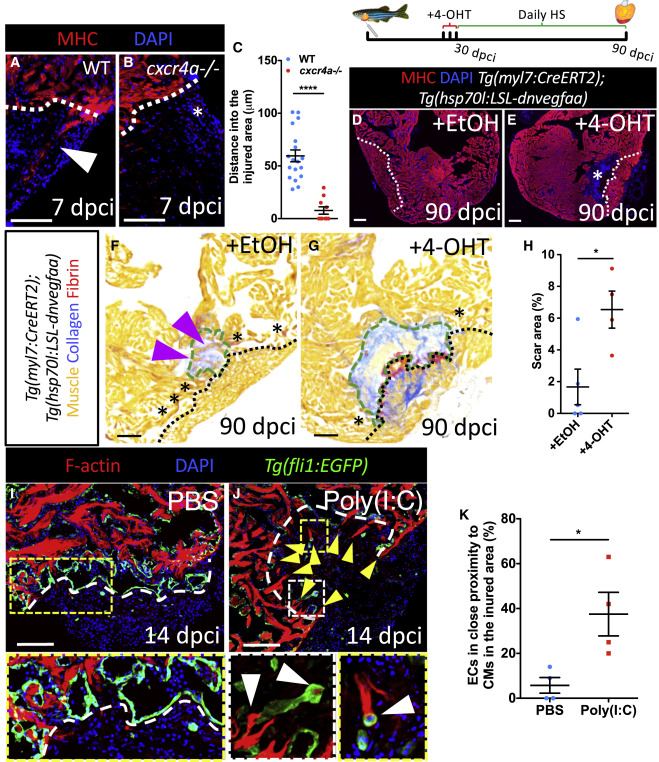
Fig. 6 Perturbations in Coronary Revascularization Impair Cardiomyocyte Repopulation (A and B) WT (A) and cxcr4a−/− (B) ventricles at 7 dpci stained for CMs (red) and DNA (blue). Presence (A) and absence (B) of cortically located regenerating CMs is indicated by white arrowhead and asterisk, respectively. (C–G) Distance spanned into the injured area by cortically located CMs in WT (n = 6) and cxcr4a−/− (n = 6) at 7 dpci. Control ([D], +EtOH) and recombined ([E], +4-OHT) ventricles at 90 dpci stained for CMs (red) and DNA (blue). Asterisk marks an area devoid of CMs (E). Control ([F], +EtOH) and recombined ([G],+4-OHT) ventricles at 90 dpci stained with AFOG to identify muscle (orange), collagen (blue), and fibrin (red). Asterisks mark contact areas between spared and regenerated myocardium. Magenta arrowheads point to regenerating CMs inside the remaining scar (F). Green dashed lines outline the scar area. (H–J) Quantification of scar area in control (n = 5) and recombined (n = 4) ventricles at 90 dpci (H). Ventricles from control ([I], PBS) and poly(I:C) (J) injected medaka at 14 dpci stained for CMs (red), DNA (blue), and endothelium (green). Yellow arrowheads point to CMs expanding beyond the border zone endocardium into the injured area; white arrowheads point to ECs in close proximity to CMs in the injured area. (K) Quantification of EC to CM association index in control (n = 4) and poly(I:C) (n = 4) injected medaka at 14 dpci. White (A, B, D, E, I, and J) and black (F and G) dotted lines delineate injured area. Data in graphs expressed as mean ± SEM. ∗p < 0.05, ∗∗∗∗p < 0.0001. Scale bars: 100 μm.
Reprinted from Developmental Cell, 51, Marín-Juez, R., El-Sammak, H., Helker, C.S.M., Kamezaki, A., Mullapuli, S.T., Bibli, S.I., Foglia, M.J., Fleming, I., Poss, K.D., Stainier, D.Y.R., Coronary Revascularization During Heart Regeneration Is Regulated by Epicardial and Endocardial Cues and Forms a Scaffold for Cardiomyocyte Repopulation, 503-515.e4, Copyright (2019) with permission from Elsevier. Full text @ Dev. Cell