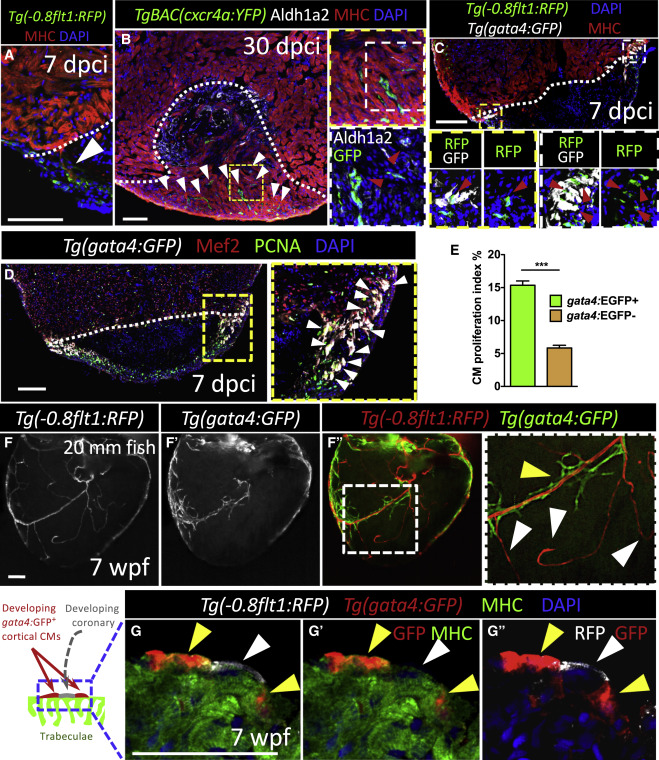
Fig. 5 Coronaries Constitute a Scaffold Available to Cardiomyocytes during Regeneration as well as during Development (A) 7 dpci ventricle stained for coronaries (green), CMs (red), and DNA (blue) (n = 8). CMs associated with regenerating coronaries in the injured area (white arrowhead). (B) 30 dpci ventricle stained for cxcr4a:YFP expression (green), activated endocardium and epicardium (white), CMs (red), and DNA (blue) (n = 6). White arrowheads point to regenerating coronaries scaffolding the regenerating myocardium. High-magnification images of regenerating intra-ventricular coronaries scaffolding the regenerating myocardium (red arrowheads). (C) 7 dpci ventricle stained for coronaries (green), gata4:GFP expression (white), DNA (blue), and CMs (red) (n = 6). High-magnification image shows regenerating gata4:GFP+ CMs associated with coronaries in the injured area (red arrowheads). (D) 7 dpci ventricle stained for gata4:GFP expression (white), CM nuclei (red), PCNA (green), and DNA (blue). High-magnification image shows cortically located and proliferating gata4:GFP+ CMs. (E) Proliferation index of gata4:GFP+ versus gata4:GFP− CMs at 7 dpci (n = 6). (F–F”) wholemount images of hearts from 20 mm long 7 wpf fish (n = 6). High-magnification images of developing coronaries (RFP+) and cortical CMs (GFP+). White arrowheads point to vessels at the sprouting front of the developing coronary network (not associated with developing cortical CMs); yellow arrowhead points to cortical CMs (associated with coronaries). (G-G”) Ventricle from a 7 wpf fish stained for coronary ECs (white), gata4:GFP expression (red), CMs (green), and DNA (blue). Yellow arrowheads point to developing cortical CMs. Coronary vessel (white arrowhead) flanked by gata4:GFP+ CMs. White dotted lines delineate injured area. Data in graph expressed as mean ± SEM. ∗∗∗p < 0.001. Scale bars: 100 μm (A–F); 50 μm (G). See also Figure S6.
Reprinted from Developmental Cell, 51, Marín-Juez, R., El-Sammak, H., Helker, C.S.M., Kamezaki, A., Mullapuli, S.T., Bibli, S.I., Foglia, M.J., Fleming, I., Poss, K.D., Stainier, D.Y.R., Coronary Revascularization During Heart Regeneration Is Regulated by Epicardial and Endocardial Cues and Forms a Scaffold for Cardiomyocyte Repopulation, 503-515.e4, Copyright (2019) with permission from Elsevier. Full text @ Dev. Cell