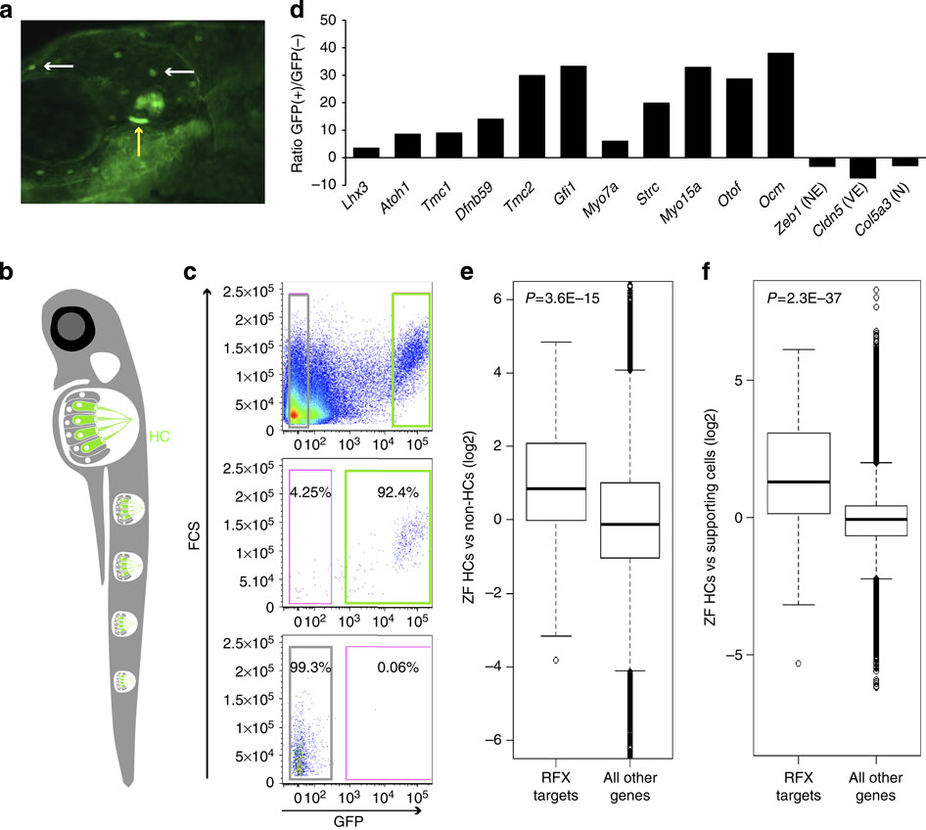
Fig. 3
The expression of RFX target genes is elevated in zebrafish HCs.
(a) An image of a 5 dpf ppv3b:GFP larvae showing GFP expression in the inner ear HCs (yellow arrow) and in the neuromast HCs (white arrows). (b) Schematic representation of a zebrafish showing the neuromast HCs. (c) Representative image of flow cytometry from dissociated 5 dpf ppv3b:GFP larvae. Top—dot plot analysis reveals a distinct population of GFP(+) cells (right). Post-sort analyses showing a cell type-specific purity >92% for GFP(+) cells (middle panel), >99% for negative cells (bottom). (d) The expression of zebrafish orthologues of known murine HC-enriched genes. Expression is enriched in the zebrafish GFP(+) cell population, while markers of NECs (for example, Zeb1), vascular epithelial cells (VE; for example, Cldn5) and neuronal cells (N; for example, Col5a3) are decreased (results based on RNA-seq). (e) Expression of the zebrafish orthologous genes of the 225 putative targets of RFX that we identified in murine HCs, was significantly elevated in zebrafish HC (GFP(+) cells). Ratios between expression level in the GFP(+) and GFP(−) cells were calculated for each gene in the zebrafish data set. The plot compares the distribution of these ratios (in log2 scale; y-axis) between the set of orthologues of the RFX targets (left boxplot) and all the other genes in the data set (right boxplot). The box indicates the first and third quartiles; the horizontal band inside the box indicates the median. The whiskers extend to the most extreme data point, which is no more than 1.5 times the interquartile range from the box. P value calculated using Wilcoxon test. (f) Similar to panel (e), but using an independent zebrafish gene expression data set. Here too, expression of RFX target genes is significantly elevated in HCs as compared with their expression levels in non-HCs.