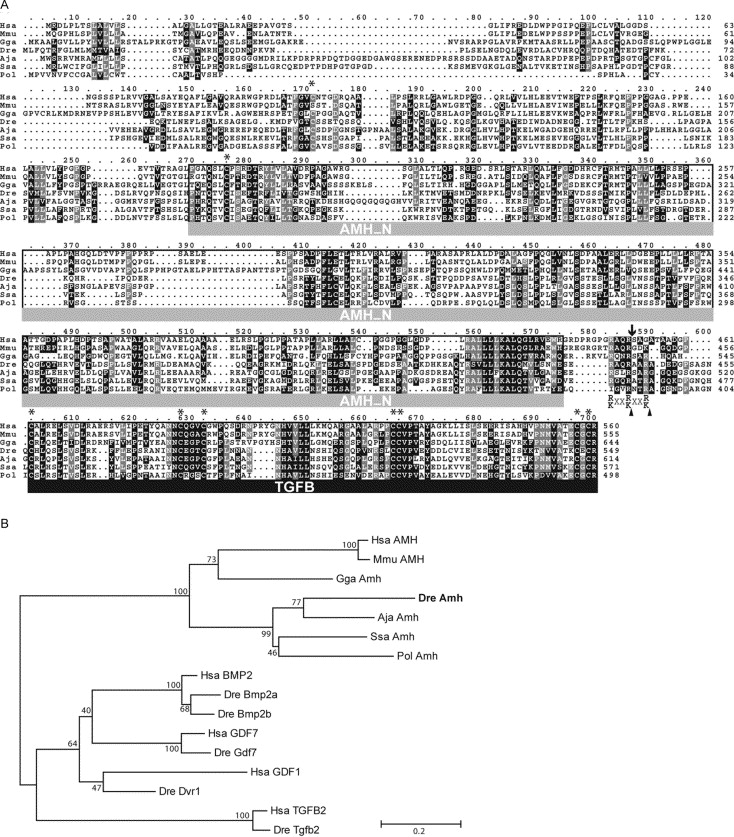
Fig. 2 Comparative analysis of zebrafish Amh. (A) Protein alignment of zebrafish Amh with other related members of the Amh family. Conserved positions with the same amino acid (black) or similar residues (grey) in more than 70% of the sequences are shadowed. The TGF-beta domain at the C-terminus (TGFB in black under bar) and the AMH domain (AMH_N in grey under bar) that characterize this protein family are labeled. Putative consensus cleaving target sites (arrowheads in R/K-XX-R/K) and the cleavage site in the human AMH (arrow) by which the active TGF-beta domain is released are indicated. Each of the eight positions with conserved cysteines is labeled with an asterisk. (B) Protein sequence relationships between the Amh family and other protein families belonging to the TGF-beta superfamily are shown in a phylogenetic tree. The branch lengths are drawn to scale to the evolutionary distance based on Poison-corrected Neighbor-Joining method, and the numbers are the percentage of bootstrap values supporting each node from 500 replicas. Scale bar, 0.2 substitutions per site. Aja, Anguilla japonica (Japanese eel); Dre, Danio rerio (zebrafish); Gga, Gallus gallus (chicken); Hsa, Homo sapiens (human); Mmu, Mus musculus (mouse); Pol, Paralichthys olivaceus (Japanese flounder); Ssa, Salmo salar (Atlantic salmon).
Reprinted from Gene expression patterns : GEP, 5(5), Rodríguez-Marí, A., Yan, Y.L., Bremiller, R.A., Wilson, C., Cañestro, C., and Postlethwait, J.H., Characterization and expression pattern of zebrafish anti-Müllerian hormone (amh) relative to sox9a, sox9b, and cyp19a1a, during gonad development, 655-667, Copyright (2005) with permission from Elsevier. Full text @ Gene Expr. Patterns