- Title
-
TNF Induces Pathogenic Programmed Macrophage Necrosis in Tuberculosis through a Mitochondrial-Lysosomal-Endoplasmic Reticulum Circuit
- Authors
- Roca, F.J., Whitworth, L.J., Redmond, S., Jones, A.A., Ramakrishnan, L.
- Source
- Full text @ Cell
|
Ceramide Causes Necrosis through Cathepsin D, BID, and BAX (A) Cartoon of TNF-mediated necrosis pathway components. CYPD, cyclophilin D; M, mitochondrion; L, lysosome. (B) Confocal images of granulomas in 3 or 5 dpi TNF-high or control larvae with yellow fluorescent macrophages infected with red fluorescent Mm. Arrowheads, extracellular bacteria; arrows, extracellular, cording bacteria. Scale bar, 100 μm. (C) Cording in 5 dpi TNF-high and control larvae. (D) Cording in 5 dpi TNF-high or control larvae treated with pepstatin A. (E) Cording in 5 dpi TNF-high or control larvae treated with E64d. (F) Cording in 5 dpi TNF-high and control larvae that are wild-type (WT) or cathepsin D morphant. (G) Cording in 5 dpi TNF-high and control larvae that are WT or BID morphant. (H) Cording in 5dpi TNF-high and control larvae treated with BI-6C9. (I) Cording in 5 dpi TNF-high or control larvae that are WT or BAXA mutant. (J) Cording in 5 dpi TNF-high and control larvae that are WT or BAXB mutant. (C–J) ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001 (Fisher’s exact test). Each panel representative of 3–6 independent experiments. See also |
|
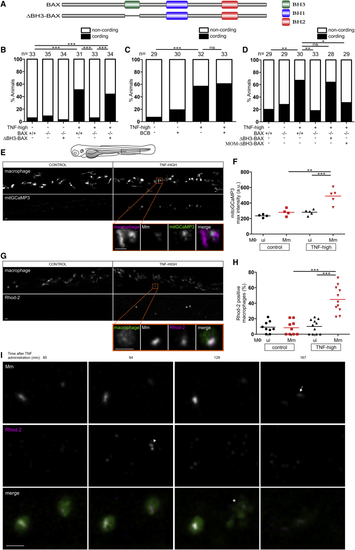
BAX Mediates Macrophage Necrosis by Promoting Mitochondrial Ca2+ Overload Independent of BH3-Dependent Oligomerization and Interaction with the Mitochondrial Outer Membrane (A) Schematic of BAX to show BH domains. (B) Cording in 5 dpi TNF-high and control larvae that are WT, BAX mutant, or BAX mutant expressing ΔBH3-BAX. ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 6 independent experiments. (C) Cording in 5 dpi TNF-high and control larvae treated with BAX channel blocker (BCB). ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 3 independent experiments. (D) Cording in 5 dpi TNF-high and control larvae that are WT, BAX mutant, BAX mutant expressing ΔBH3-BAX, or BAX mutant expressing ΔBH3-BAX targeted to the mitochondrial outer membrane (MOM-ΔBH3-BAX). ∗p < 0.05; ∗∗p < 0.01 (Fisher’s exact test). Representative of 2 independent experiments. (E) Representative confocal images of 1 dpi TNF-high (90 min post-TNF treatment) or control larvae, both expressing mitGCaMP3 (GCaMP3 targeted to the mitochondria) in their macrophages, which are red fluorescent. Area shown corresponds to area in the shaded rectangle in the cartoon above. Detailed image of orange rectangle in TNF-high larva additionally shows the bacteria within a mitGCaMP3-expressing macrophage. Scale bars, 10 μm. (F) Quantitation of mitGCaMP3 fluorescence in individual macrophages from larvae in (A). Black and red symbols represent uninfected (ui) and Mm-infected macrophages, respectively, in the same control or TNF-treated animal. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (G) Representative confocal images of 1 dpi TNF-high (120 min post-TNF treatment) or control larvae with yellow fluorescent macrophages showing Rhod-2 fluorescence (red). Area shown similar to (A). Detail: macrophage in the orange rectangle. Scale bar, 10 μm. (H) Percentage of uninfected or Mm-infected macrophages with Rhod-2 fluorescence from TNF-high larvae in (C). Black and red symbols represent uninfected and Mm-infected macrophages, respectively, in the same control or TNF-treated animal. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (I) Time-lapse confocal images of two infected macrophages in a 1 dpi larva at indicated time points after TNF administration. Arrowhead, Rhod-2 positive macrophage; asterisk, dead macrophage; arrow, extracellular bacteria. Scale bar, 10 μm. See also |
|
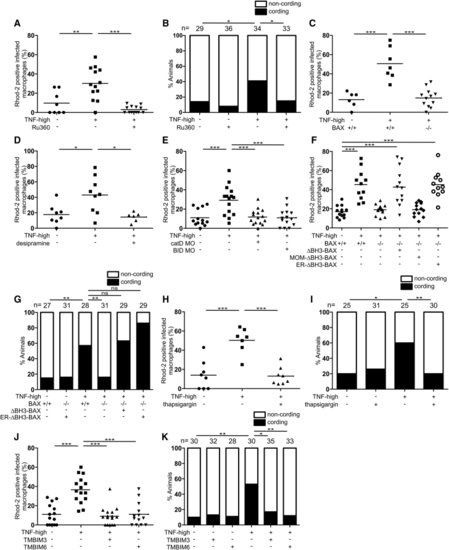
BAX Promotes Ca2+ Flow from the ER into the Mitochondrion (A) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with Ru360. Horizontal bars, means; ∗∗p < 0.01; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (B) Cording in 5 dpi TNF-high and control larvae treated with Ru360. ∗p < 0.05 (Fisher’s exact test). Representative of 5 independent experiments. (C) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae that are WT or BAX mutant. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (D) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with desipramine. Horizontal bars, means; ∗p < 0.05 (one-way ANOVA with Tukey’s post-test). (E) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae that are WT, cathepsin D, or BID morphant. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (F) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae that are WT, BAX mutant, or BAX mutant expressing ΔBH3-BAX, MOM- or ER-targeted ΔBH3-BAX. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (G) Cording in 5 dpi TNF-high or control larvae that are WT, BAX mutant, BAX mutant expressing ΔBH3-BAX, or BAX mutant expressing ER-targeted ΔBH3-BAX. ∗∗p < 0.01 (Fisher’s exact test). Representative of 2 independent experiments. (H) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with thapsigargin. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (I) Cording in 5 dpi TNF-high or control larvae treated with thapsigargin. ∗p < 0.05; ∗∗p < 0.01 (Fisher’s exact test). Representative of 3 independent experiments. (J) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae that are WT or overexpressing TMBIM3 or TMBIM6. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (K) Cording in 5 dpi TNF-high or control larvae that are WT or overexpressing TMBIM3 or TMBIM6. ∗p < 0.05; ∗∗p < 0.01 (Fisher’s exact test). Representative of 2 independent experiments. |
|
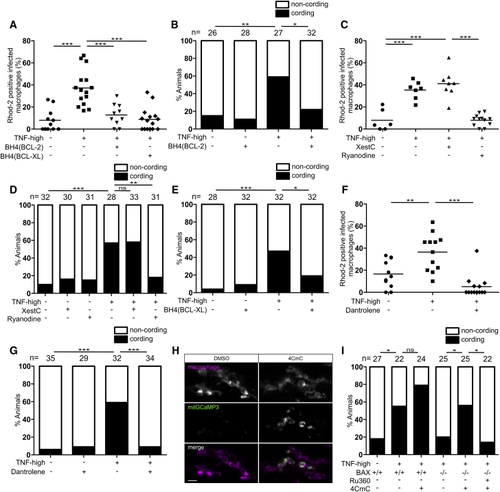
Ryanodine Receptor Mediates Macrophage Necrosis in Excess TNF Conditions (A) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae that are WT or expressing the BH4 domain of BCL-2 or BCL-XL. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (B) Cording in 5 dpi TNF-high and control larvae that are WT or expressing the BH4 domain of BCL-2. ∗p < 0.05; ∗∗p < 0.01 (Fisher’s exact test). Representative of 3 independent experiments. (C) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with xestospongin C (XestC) or ryanodine. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (D) Cording in 5 dpi TNF-high and control larvae treated with XestC or ryanodine. ∗∗p < 0.01; ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 2 independent experiments. (E) Percentage of larvae with cording among WT or siblings expressing the BH4 domain of BCL-XL injected with TNF. ∗p < 0.05; ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 2 independent experiments. (F) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with dantrolene. Horizontal bars, means; ∗∗p < 0.01; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (G) Cording in 5 dpi TNF-high and control larvae treated with dantrolene. ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 4 independent experiments. (H) Representative confocal images of 3 day larvae expressing GCaMP3 targeted to the mitochondria of their macrophages which are red fluorescent, 2 h after administration of 4CmC larvae. Scale bar, 10 μm. (I) Cording in 5 dpi TNF-high and control larvae that are WT or BAX mutant and treated with Ru360 and/or 4CmC. ∗p < 0.05 (Fisher’s exact test). Representative of 2 independent experiments. See also |
|
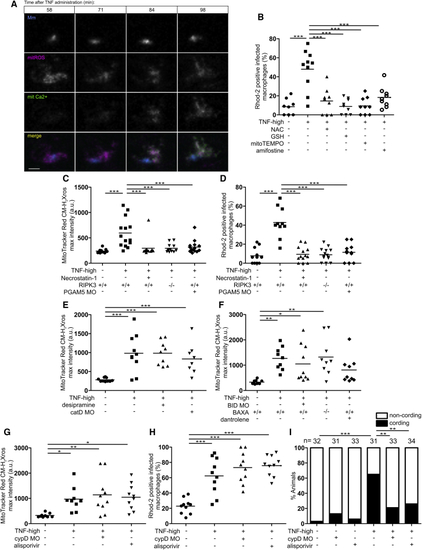
Mitochondrial ROS Launch the Pathogenic Mitochondrial-Lysosomal-ER Circuit that Leads to Mitochondrial Ca2+ Overload and Macrophage Necrosis (A) Time-lapse confocal images of an infected macrophage in a 1 dpi mitGCaMP3 larva at indicated time points after TNF administration with MitoTracker Red CM-H2Xros. Scale bar, 5 μm. (B) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae treated with NAC, GSH, MitoTEMPO, or amifostine. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (C) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae that are WT, WT treated with necrostatin-1, PGAM5 morphant, or RIPK3 mutant. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (D) Percentage of Rhod-2-positive macrophages in 1 dpi control and TNF-high larvae control, that are WT, WT treated with necrostatin-1, PGAM5 morphant, or RIPK3 mutant. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (E) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae that are WT, WT treated with desipramine, or cathepsin D morphant. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (F) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae that are WT, WT treated with dantrolene, BID morphant, or BAX mutant. Horizontal bars, means; ∗p < 0.05; ∗∗p < 0.01 (one-way ANOVA with Tukey’s post-test). (G) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae that are WT, WT treated with alisporivir, or cyclophilin D morphant. Horizontal bars, means; ∗p < 0.05; ∗∗p < 0.01 (one-way ANOVA with Tukey’s post-test). (H) Percentage of Rhod-2-positive macrophages in 1 dpi TNF-high or control larvae that are WT, WT treated with alisporivir, or cyclophilin D morphant. Horizontal bars, means; ∗∗∗p < 0.001 (Fisher’s exact test). (I) Cording in 5 dpi TNF-high or control larvae that are WT, WT treated with alisporivir, or cyclophilin D morphant. ∗∗p < 0.01; ∗∗∗p < 0.001 (Fisher’s exact test). See also |
|
Pharmacological Inhibition of Voltage-Gated L-Type Ca2+ Channels Inhibits TNF-Mediated Macrophage Mitochondrial Ca2+ Overload and Necrosis in Mm- and Mtb-Infected Larvae (A) Percentage of Rhod-2-positive macrophages in 1 dpi control or TNF-high larvae treated with nifedipine, verapamil, or diltiazem. Horizontal bars, means; ∗∗p < 0.01; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). Representative of 2 independent experiments. (B–D) Cording in 5 dpi control or TNF-high larvae treated with nifedipine (B), diltiazem (C), and verapamil (D). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001 (Fisher’s exact test). Representative of 2–4 independent experiments. (E) Representative confocal images of 1 dpi larvae with yellow fluorescent macrophages infected with 100 WT Mm or 80 Mtb (both red fluorescent). Scale bar, 50 μm. (F) Quantitation of mitochondrial ROS production in infected macrophages of 1 dpi TNF-high or control larvae infected with 100 WT Mm or 80 Mtb. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (G) Quantitation of mitGCaMP3 fluorescence in infected macrophages in 1 dpi TNF-high or control larvae infected with 100 WT Mm or 185–200 Mtb treated with dantrolene, nifedipine, diltiazem, or verapamil. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). |
|
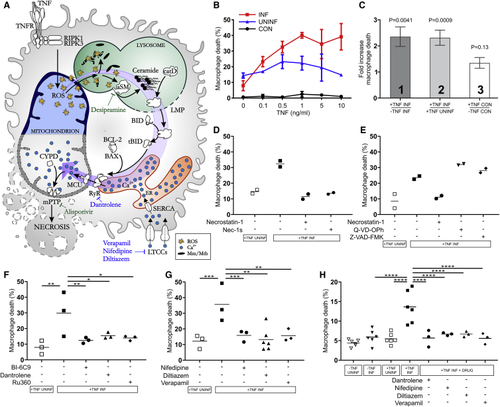
Mycobacterium-Infected Human Macrophages Undergo TNF-Mediated Necrosis (A) Model of TNF-mediated macrophage necrosis pathway. LMP, lysosomal membrane permeabilization; previously identified drugs in green, drugs identified in this study in blue. (B) THP-1 macrophage death 5 h after TNF administration (Mean ± SD). CON, macrophages from uninfected wells; INF, Mm-infected macrophages in infected wells; UNINF, uninfected macrophages in infected wells. (C) Quantification of TNF-induced macrophage death in multiple experiments. Column 1: ratio of TNF-treated to vehicle-treated dead Mm-infected macrophages. Column 2: ratio of dead infected macrophages to dead uninfected macrophages in the same TNF-treated well. Column 3: ratio of dead macrophages in TNF-treated to vehicle-treated uninfected wells. Mean ± SEM of 12 independent experiments for columns 1 and 2, and 11 independent experiments for column 3; one sample t test to a hypothetical value of 1, corresponding to the null hypothesis that TNF and infection do not influence cell death. (D) Percentage of dead Mm-infected macrophages after TNF administration treated with necrostatin-1 or Nec-1 s. Horizontal bars, means. Representative of 3 independent experiments. (E) Percentage of dead Mm-infected macrophages after TNF administration treated with necrostatin-1, Q-VD-OPh, or Z-VAD-FMK. Horizontal bars, means. Representative of 3 (necrostatin-1 and Z-VAD-FMK) or 2 (Q-VD-OPh) independent experiments. (F and G) Percentage of dead Mm-infected macrophages after TNF administration treated with BI-6C9, dantrolene, Ru360 (F), and diltiazem, nifedipine, or verapamil (G). Horizontal bars, means; ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001 (one-way ANOVA with Bonferroni’s post-test for comparisons shown). (H) Percentage of dead Mtb-infected macrophages after TNF administration treated with dantrolene, diltiazem, nifedipine, or verapamil. Horizontal bars, means; ∗∗∗∗p < 0.0001 (one-way ANOVA with Bonferroni’s post-test for comparisons shown). See also |
|
Camptothecin-Induced Apoptosis Is Decreased in BAXA and BAXB Mutant Zebrafish, Related to (A) Global analysis without end-gap penalty of protein sequence homology between human BAK1 (transcripts 202 and 203), BAX (only major transcripts, alpha and beta) and zebrafish BAXA and BAXB. Prefix h, human; prefix zf, zebrafish. Although BAXB has been suggested to be the zebrafish functional equivalent of human BAK ( (B) Comparison of protein sequence homology between human BAX alpha and zebrafish BAXA. Relevant BH domains of BAX are showed in colored boxes. (C) Representative inverted fluorescence images of 2 dpf larvae that are WT, BAXA mutant or BAXB mutant treated with camptothecin and incubated with acridine orange to detect apoptotic cells. (D) Quantification of camptothecin-induced apoptosis (See |
|
TNF Triggers Mitochondrial ROS Production Only in Infected Macrophages, Related to (A) Representative confocal images of 1 dpi TNF-high or control larvae with yellow fluorescent macrophages, showing MitoTracker Red CM-H2Xros fluorescence corresponding to similar area of the fish as in (B) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae. Each point represents the mean of maximum intensity fluorescence of MitoTracker Red CM-H2Xros per fish from images in (A). Black and red symbols represent uninfected and Mm-infected macrophages, respectively, in the same control or TNF-administered animal. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). (C) Representative confocal images of 1 dpi TNF-high or control larvae showing MitoSOX fluorescence (red) corresponding to similar area of the fish as in (D) Quantification of mitochondrial ROS production in 1 dpi TNF-high or control larvae. Each point represents the mean of maximum intensity fluorescence of MitoSOX per fish from images in (C). Black and red symbols represent uninfected and Mm-infected macrophages, respectively, in the same control or TNF-administered animal. Horizontal bars, means; ∗∗∗p < 0.001 (one-way ANOVA with Tukey’s post-test). |
Reprinted from Cell, 178(6), Roca, F.J., Whitworth, L.J., Redmond, S., Jones, A.A., Ramakrishnan, L., TNF Induces Pathogenic Programmed Macrophage Necrosis in Tuberculosis through a Mitochondrial-Lysosomal-Endoplasmic Reticulum Circuit, 1344-1361.e11, Copyright (2019) with permission from Elsevier. Full text @ Cell