- Title
-
In vivo single-cell lineage tracing in zebrafish using high-resolution infrared laser-mediated gene induction microscopy
- Authors
- He, S., Tian, Y., Feng, S., Wu, Y., Shen, X., Chen, K., He, Y., Sun, Q., Li, X., Xu, J., Wen, Z., Qu, J.Y.
- Source
- Full text @ Elife
|
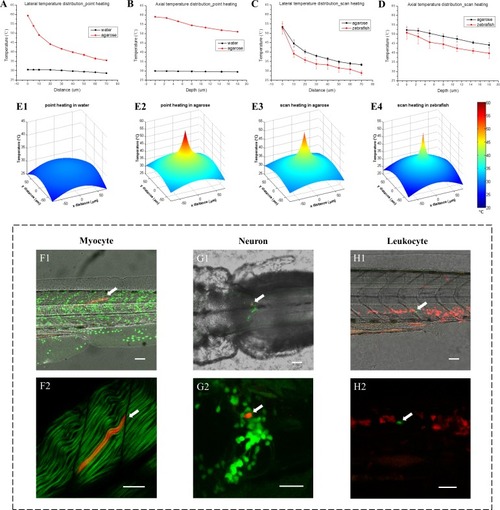
3D temperature distribution measured through fluorescent thermometry and single-cell gene induction in zebrafish.(A) Lateral temperature distributions with IR laser heating in water and 3% agarose (95 mW IR laser was focused in samples for point heating). (B) Corresponding axial temperature distributions in water and 3% agarose. (C) Lateral temperature distributions with IR laser scan heating in 3% agarose and zebrafish muscle in vivo (95 mW IR laser was scanned in an 8 µm × 8 µm region during heating to avoid tissue injury). (D) Corresponding axial temperature distributions in 3% agarose and zebrafish muscle in vivo. Each statistical distribution curve in (A–D) is shown in terms of the mean with the standard deviation over more than five measurements. (E1–E4) 3D view of lateral temperature distributions with IR laser point and scan heating in water, 3% agarose and zebrafish muscle, respectively. (F1) A merged image of bright-field (gray) middle trunk, Dendra2-labeled nuclei (green) and a single myocyte expressing DsRedx via heat shock gene induction (red) in a one dpf zebrafish. (F2) A merged image of second harmonic generation (SHG) of muscle fibers (green) and the single myocyte expressing DsRedx (red). (G1) A merged image of bright-field (gray) hindbrain, GFP-labeled tyrosine hydroxylase-positive (th-positive) neurons (green) and a single neuron expressing DsRedx via heat shock gene induction (red) in a three dpf zebrafish. (G2) An enlarged image of (G1) with GFP and DsRedx-labeled neurons (by maximum projections). (H1) A merged image of bright-field (gray) middle trunk, DsRedx-labeled leukocytes (red) and a single cell expressing GFP via heat shock gene induction (green) in a two dpf zebrafish. (H2) An enlarged image of (H1) with merged DsRedx and GFP-labeled leukocytes. Arrows in (F–H): heat-shock labeled single myocyte, neuron and leukocyte, respectively. Scale bars: 50 µm (F1–H1); 30 µm (F2–H2). |
|
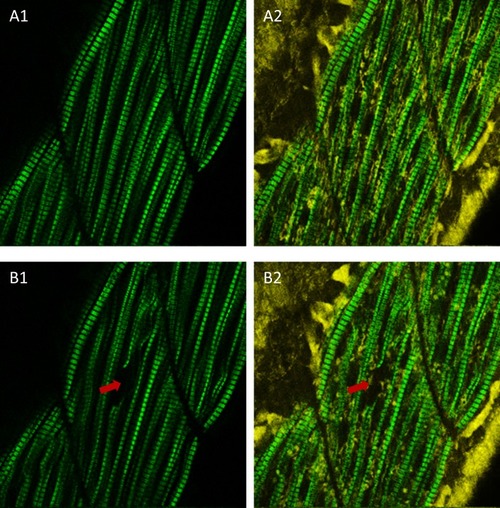
Laser-induced injury in zebrafish muscle.(A1) SHG image of zebrafish muscle before laser heating. (A2) Merged image of SHG and reduced nicotinamide-adenine dinucleotide (NADH) autofluorescence signals of zebrafish muscle before laser heating. (B1) SHG image of zebrafish muscle after focusing 85 mW IR laser to a single fiber for heating over 3 s. (B2) Merged image of SHG and NADH autofluorescence signals of zebrafish muscle after IR laser heating. The arrows in (B) indicate the laser heating point, where the sarcomere structure disappeared, indicating that muscle fiber was injured. To optimize the heat shock gene induction and avoid cell damage, scan heating should be performed for in vivo heat shock in zebrafish. The field of view of (A) and (B): 100 µm × 100 µm. |
|
The high-precision IR-LEGO-mediated single HE lineage tracing.(A) The genetic labeling diagram of the lineage tracing experiment. The heat shock-induced CreERT2 line Tg(hsp70l:mCherry-T2a-CreERT2) is crossed with the double reporter fish Tg(kdrl:loxP-DsRedx-loxP-EGFP;coro1a:loxP-DsRedx-loxP-EGFP) to obtain triple transgenic Tg(kdrl:loxP-DsRedx-loxP-EGFP; coro1a:loxP-DsRedx-loxP-EGFP; hsp70l:mCherry-T2a-CreERT2) embryo. Upon IR laser illumination, the targeted HEs will express CreER. After 4-OHT treatment, the CreER will enter the cell nucleus and remove the DsRedx cassettes flanked by loxP from the genome. After that, the target HEs and their hematopoietic progenies will express GFP signal, thus be distinguished from the unlabeled DsRedx+cells. (B) The work flow of the IR-LEGO-mediated single HE lineage tracing assay. (C) Representative time-lapse images of the single-HE heat-shocked embryos (upper row) and control embryos (lower row). Dotted lines depict the caudal aorta (CA) in the PBI region (dorsal wall on the top). Number at the top left corner in each image indicates the developmental stage of the embryos (hh:mm post fertilization). The heat-shock labeled HE (white arrow) on the ventral wall of CA gradually turns on the expression of GFP without affecting neighbor HEs, while the HEs in control embryos do not express GFP during time-lapse imaging. (D) Images of GFP+ T cells and myeloid cells in single-HE labeled fish and control fish at 7 dpf (back on the top). The left column shows small and round coro1a:GFP+ T cells in the thymus (depicted by dashed lines). The right column shows coro1a:GFP+ myeloid cells on the trunk, which have irregular shape. While GFP+ T cells and myeloid cells are persistently observed in many of the labeled fish (upper row), rare GFP signals are detected in most of the control fish (lower row). (E) Quantification of GFP+ T cells and myeloid cells in both single HE-labeled fish (n=27) and control fish (n=109) at 7 dpf. Statistical analysis indicates that for both T cells and myeloid cells, the GFP+ cell number in heat-shock labeled fish is significantly higher than that in control fish. The cell numbers are shown in terms of mean ± standard error of the mean. The Mann–Whitney–Wilcoxon rank-sum test was used for significance test. ***P < 0.001. |
|
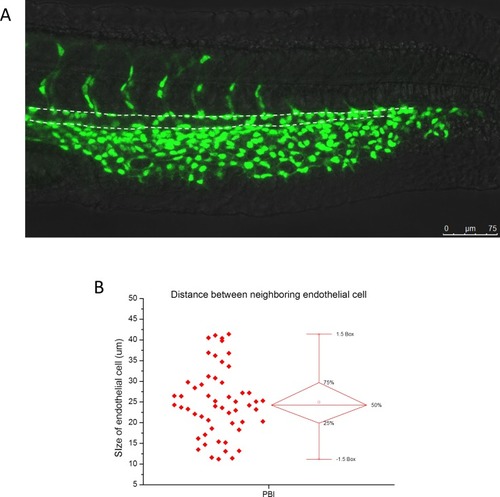
Size of endothelial cells on the ventral wall of aorta in the PBI region.(A) The representative fluorescence image of a 28 hpf Tg(kdrl:GFP; kdrl:nlsEOS) zebrafish with endothelial nuclei labeled by nls-EOS (green). The region marked by dashed lines indicates aorta in the PBI. (B) Statistics of the distance between the nuclei of neighboring endothelial cells on the aortic floor (bottom dashed line in (A)) in the PBI region. The average distance is 24.9 µm and the standard deviation is 8.2 µm. The total number of endothelial cells is 54 from three zebrafish. |
|
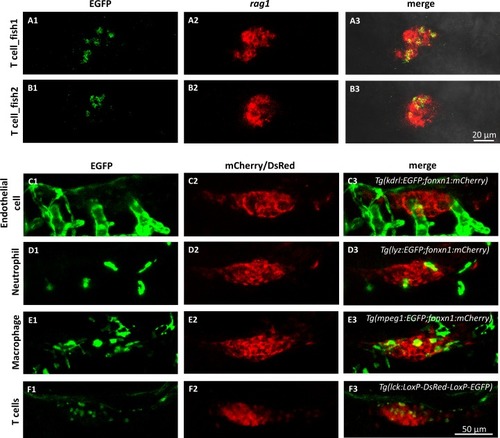
Verification of T cells in the thymus of the single-HE labeled zebrafish at 7dpf.(A1 and B1) The representative images (by maximum projections) of antibody stained EGFP+ cells in the thymus of triple Tg zebrafish. (A2 and B2) The patterns of rag1 expression in the thymus (by maximum projections) revealed by whole-mount in situ hybridization. (A3 and B3) The colocalization of EGFP with rag1 expression indicates that the EGFP+ cells in the thymus of 7dpf zebrafish are T cells. In addition, T cells (F) can be distinguished from other cell types (C–E) in the thymus based on their small and round shapes. (C) In vivo dual-color imaging of blood vessel endothelium (EGFP) and thymus epithlium (mCherry). (D) In vivo dual-color imaging of neutrophils (EGFP) and thymus epithlium (mCherry). (E) In vivo dual-color imaging of macrophages (EGFP) and thymus epithlium (mCherry). (F) In vivo dual-color imaging of heat shocked T cells (EGFP) and non-heat shocked T cells in thymus (DsRed). |
|
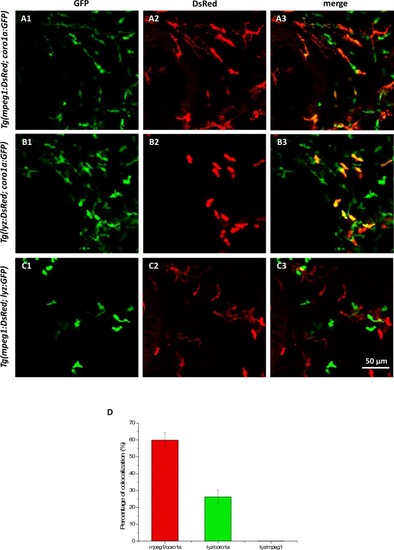
Verification of myeloid cells on the trunk of the single-HE labeled zebrafish at 7dpf.(A1–A3) In vivo dual-color imaging of leukocytes (coro1a:GFP) and macrophages (mpeg1:DsRed). (B1–B3) In vivo dual-color imaging of leukocytes (coro1a:GFP) and neutrophils (lyz:DsRed). (C1–C3) In vivo dual-color imaging of neutrophils (lyz:GFP) and macrophages (mpeg1:DsRed). (D) The colocalization percentages of mpeg1+/coro1a+ and lyz+/coro1a+ cells are 59.9% and 26.3%, respectively (n = 3), while lyz+ cells are not colocalized with mpeg+ cells, indicating that the majority of coro1a+ cells (86.2%) on the trunk are lyz+ or mpeg1+ myeloid cells. |
|
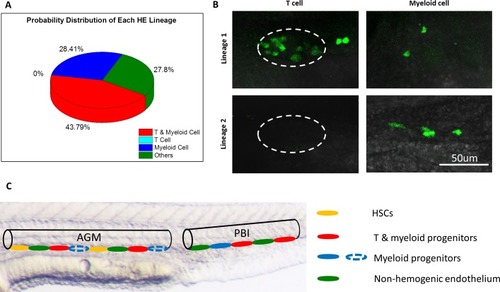
The heterogeneous hematopoietic lineages of HE.(A) Distribution of probability of each HE subpopulation. At least two subpopulations of HEs with distinct hematopoietic potentials exist in the PBI. One group of HEs has lymphoid and myeloid potential, while the other group generates myeloid lineage only. HEs with lymphoid lineage potential only were not found. The total number of single-HE labeled zebrafish is 27. (B) Representative images of GFP+ T cells and myeloid cells derived from two distinct HE subpopulations. It shows that the myeloid-lymphoid bipotent HEs give rise to both T cells and myeloid cells (the upper row), while the myeloid unipotent HEs generate myeloid progenies exclusively (the lower row). The left column shows small and round coro1a:GFP+ T cells in the thymus (depicted by dashed lines). The right column shows coro1a:GFP+ myeloid cells on the trunk, which have irregular shape. (C) A schematic diagram illustrates the heterogeneity of hematopoietic potential of HEs. The aortic lumen in AGM and PBI are represented by black lines. The different HE subpopulations on the ventral floor of aorta are indicated by ellipses with different colors. The HEs in PBI give rise to T lymphoid-myeloid bipotent progenitors (red) and myeloid unipotent progenitors (blue), while the HEs in the AGM produce HSCs (orange) and T lymphoid-myeloid bipotent progenitors. It is possible that the AGM HEs can also generate myeloid unipotent progenitors (blue dotted ellipses), similar to their counterparts in the PBI. |
|
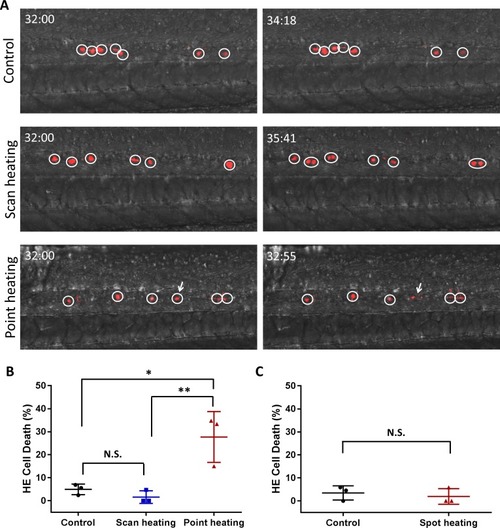
Assessment of HE cell damage caused by IR-LEGO heat shock.(A) Representative time-lapse images of the control embryos (the first row), the scanning heat-shocked embryos (the second row), and the point heat-shocked embryos (the third row). Circles depict the individual converted HEs in the PBI region (dorsal wall on the bottom). Number at the top left corner in each image indicates the developmental stage of the embryos (hh:mm post fertilization). One of the point heat-shock HE (white arrow) burst into fragment, while the HEs in control embryos and scanning heat-shocked keep viability during time-lapse imaging. (B) Quantification of the HE cell death in the control embryos (three independent experiments, in which the number of dead cell/total cell are 1/18, 1/42 and 2/29, respectively. Totally 4 of 89 cells died.), the scanning heat-shocked embryos (three independent experiments, in which the number of dead cell/total cell are 1/21, 0/34 and 0/23, respectively. Totally 1 of 78 cells died.), and the point heat-shocked embryos (three independent experiments, in which the number of dead cell/total cell are 8/23, 9/27 and 3/20, respectively. Totally 20 of 70 cells died.). Statistical analysis indicates that the percentage of HE cell death in the scanning heat-shocked embryos has no difference from the control group, while heat shock fixed on a single point result in significantly more cell death. The percentages of cell death are shown in terms of mean ± standard deviation. One-way ANOVA was used for significance test. *P < 0.05; **P < 0.01. (C) Cell death test on the old IR-LEGO system used for bulk labeling previously. The IR laser beam was loosely focused by a low-NA lens and heat shocked the embryos on each spot for 2 minutes. Statistical analysis shows that the single spot heat shock used for bulk labeling would not cause cell damage. |