- Title
-
The role of Sox6 in zebrafish muscle fiber type specification
- Authors
- Jackson, H.E., Ono, Y., Wang, X., Elworthy, S., Cunliffe, V.T., Ingham, P.W.
- Source
- Full text @ Skelet Muscle
|
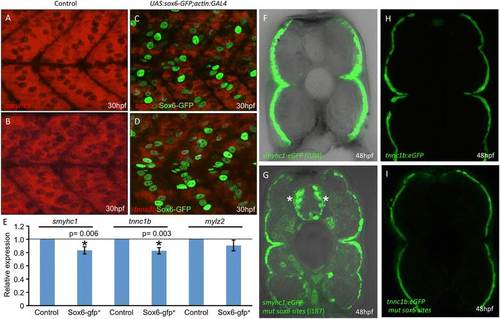
Control of slow-twitch specific gene expression by Sox6.(A,B) Fluorescent in situ hybridization of 30 hours postfertilization (hpf) wild-type embryos revealing accumulation of smyhc1 and tnnc1b transcripts throughout the mononucleated slow-twitch muscle; (C,D) transgenic embryos expressing Sox6-GFP in slow-twitch fibers (green nuclei) hybridized under identical conditions to the wild-type controls (A,B) showing reduced levels of smyhc1 and tnnc1b transcripts; (E) graphical representation of data pooled from three independent qPCR analyses of smyhc1, tnnc1b and mylz2 transcription in control and UAS:sox6-GFP;actin:GAL4 (sox6-gfp+) embryos at 30 hpf. Student’s two-tailed, unpaired t-test, error bars represent standard deviation, control n = 3, sox6-gfp+ n = 3, asterisks indicate significance; (F) Cross section through a trunk somite of a 48 hpf smyhc1:GFP transgenic embryo; note that expression is restricted to the superficially located slow-twitch muscle fibers; (G) similar preparation of an embryo transgenic for the same smyhc1:GFP reporter gene in which the putative Sox6 binding sites have been mutated: note the ectopic expression in many fast-twitch muscle fibers as well as in prominent clusters of neurons in the neural tube (asterisks). (H) Cross section through a trunk somite of a 48 hpf tnnc1b:GFP transgenic embryo: expression is similarly restricted to the slow-twitch fibers. (I) Similar preparation of a 48 hpf embryo transgenic for the same reporter in which the putative Sox6 binding sites have been mutated: note the absence of ectopic expression of this transgene. |
|
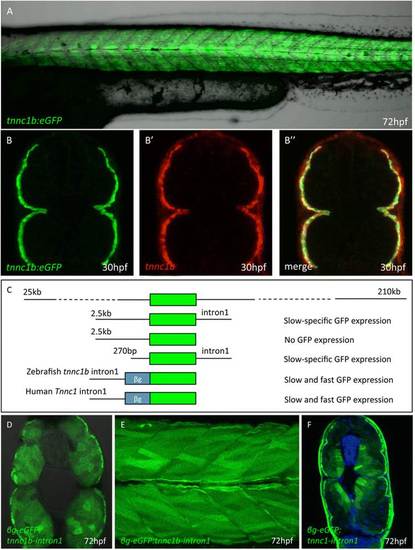
Defining thetnnc1bcis regulatory region. (A) Trunk region of a Tg(BACtnnc1b:eGFP)i293 larva 72 hpf showing reporter gene expression specifically in the slow-twitch muscle. (B) Cross section through the mid-trunk region of a 30 hpf Tg(BACtnnc1b:eGFP)i293 embryo showing reporter gene expression restricted to the superficial layer of muscle fibers: (B′) the same section showing endogenous tnnc1b mRNA assayed by fluorescent in situ hybridization. (B′′) Merged images showing that the reporter faithfully recapitulates the expression pattern of the endogenous gene. (C) Schematic representation of deletion derivatives of the BACtnnc1b:eGFP reporter construct (Green box = eGFP); the expression patterns observed in transgenic lines carrying these construct is indicated on the right. (D) Cross section and (E) optical sagittal section of a 72 hpf transgenic larva showing expression of eGFP throughout the myotome driven by the first intron of the zebrafish tnn1cb intron and a mammalian βglobin minimal promoter. (F) Cross section of a 72 hpf transgenic embryo in which expression of eGFP is driven by the first intron of the human slow troponin c gene. The strong signal in the skin in is a staining artefact and was not present in live embryos. EXPRESSION / LABELING:
|
|
Targeted mutagenesis of the zebrafishsox6locus. (A) Schematic representation of the zinc-finger nuclease designed to target genomic sequences in exon 8, upstream of the HMG box, of the zebrafish sox6 gene. (B-E) Immunohistochemical detection of Sox6 protein using a Sox6-specific antibody; Sox6 protein is detectable in the nuclei of fast muscle fibers (arrows) (B and B′) and the optic tectum (arrow) (C and C′) of wild-type embryos. In sox6 homozygous mutants, by contrast, no signal is detected in either the muscle (D and D′) or the optic tectum (E and E′), indicating the successful generation of a null mutant. (Blue signal = DAPI). |
|
tnnc1bis ectopically expressed in fast-twitch fibers of homozygoussox6mutants. Endogenous tnnc1b expression (A) and tnnc1b:eGFP reporter expression (green) (B) is restricted to slow-twitch fibers of wild-type embryos at 48 hours postfertilization (hpf) and excluded from fast-twitch fibers (B′:red) as shown in merged image (B′′). In sox6 mutants, endogenous tnnc1b(C) and the tnnc1b:eGFP reporter (D) are expressed in both slow and fast-twitch fibers (D′: red) as shown in merged image (D′′). Expression of ryr1a is restricted to slow-twitch fibers in wild type (E) but ectopically expressed in the fast-twitch fibers in sox6 mutant embryos at 30 hpf (I). By contrast, myh7b expression remains restricted to slow-twitch fibers in sox6 mutants (cf. F and J). Slow-twitch specific expression of the endogenous smyhc1(G) and the smyhc1:GFP reporter (H) in wild-type embryos. In sox6 mutants (K,L), expression is largely similarly restricted but ectopically expressed in a few fast fibers (arrows). |
|
Partial de-repression of slow-twitch genes insox6mutant larvae. (A-C)smych1, tpm2 and mylz10 remain restricted to the slow-twitch fibers in wild-type larvae at 3 or 4 days postfertilization (dpf). (D-F) All three genes are ectopically expressed in some fast-twitch fibers in sox6 mutants by 3 or 4 dpf. (G-G′′)smyhc1:GFP expression (green) is excluded from fast-twitch fibers (red) in 5 dpf wild-type larvae. (H-H′′) In sox6 mutant larvae at 5 dpf, a subset of fast-twitch fibers ectopically express the reporter gene (H). The expression of the fast-twitch myosin revealed by mAb F310 (red) in the fast domain remains unaffected in sox6 mutants (cf G′ and H′). EXPRESSION / LABELING:
PHENOTYPE:
|
|
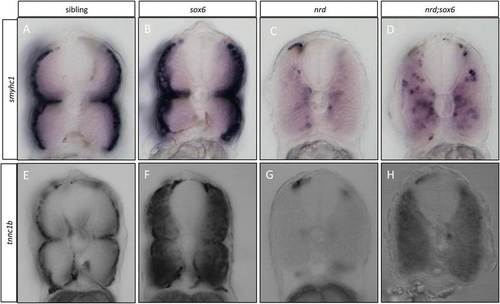
Elimination ofsox6rescuestnnc1bandsmyhc1expression inprdm1anrdmutants. (A-D) Cross sections through the trunk region of embryos that were hybridized with the smyhc1 probe at 30 hours postfertilization (hpf): note that migration of the slow-twitch progenitors is disrupted in prdm1a mutants, and the expression of smyhc1 is dramatically downregulated (cf A and C); in the prdm1a;sox6 double mutant, smyhc1 expression is restored in the adaxial cells, though their migration still appears disrupted (cf. C and D). (E-H) Similar cross sections of wild-type and mutant embryos hybridized with the tnnc1b probe. Expression of tnnc1b is completely lost prdm1a mutants (cf E and G); expression is restored throughout the myotomes in the prdm1a;sox6 double mutants, though at lower levels than in the sox6 single mutant (cf. F and H) are slow specific genes, which are expressed in the superficial slow-twitch fibers in wild-type embryos at 30 hpf (A and B). |
|
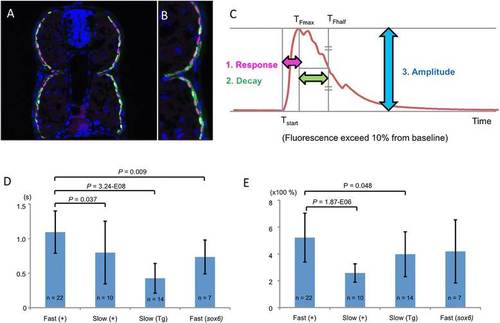
Sox6 controls the Ca2+ response of fast-twitch fibers. (A,B) Cross section through a trunk somite of a 48 hours postfertilization (hpf) Tg(smyhc1:GCaMP3)i280 embryo, showing expression of GCaMP3 (green) is restricted to slow-twitch fibers, revealed by mAbF59 labeling (red). Nuclei are stained with DAPI (blue). (C) Stereotypical representation of the GCaMP3 signal in response to chemical stimulation indicating the three parameters measured in this analysis. (D) Response time in seconds (s) and (E) amplitude of Ca2+ response in fast and slow-twitch fibers of Tg(smyhc1:GCaMP3)i280 (Tg) and transient transgenic wild-type (+) and sox6 mutant larvae (52 to 55 hpf); Student’s two-tailed, unpaired t-test; Error bars represent standard deviation. |
|
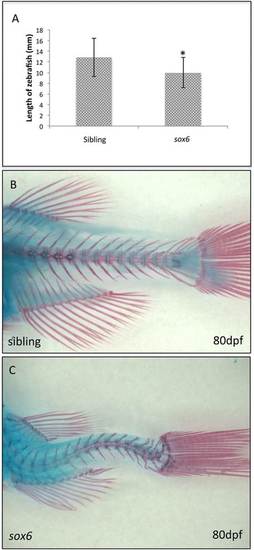
sox6mutant adults are small and have spinal deformities. (A)sox6 mutant fish are significantly smaller than siblings by 1 month. Student’s two-tailed, unpaired t-test P <0.05, error bars represent standard deviation, sibling n = 17, mutant n = 17, asterisk indicates significance. (B and C) By 80 dpf, there is clear curvature of the spine in all surviving homozygous sox6 mutant fish (C) as compared to siblings (B). PHENOTYPE:
|
|
Disruption of fiber type identity in adultsox6mutant fish. (A,B) at 3 months, expression of the tnnc1b:eGFP reporter is restricted to the medio-lateral wedge of slow-twitch fibers in wild-type fish. (C,D) In sox6 mutants, the reporter gene is also expressed at lower levels in fast-twitch fibers while the slow-twitch fibers extend closer to the midline. The diameter of the slow-twitch fibers is significantly reduced compared to wild-type (cf. B and D). (E) The smyhc1:GFP reporter is expressed at high levels in the slow-twitch fiber of 2-month-old fish. (F) the reporter shows a similar pattern of expression in sox6 mutants at the same stage, with lower level expression extending more medially than in wild type. EXPRESSION / LABELING:
PHENOTYPE:
|
|
The expression of slow-specific genes is downregulated in actin:GAL4; UAS:sox6-GFP embryos. The slow-twitch specific expression of tnnt1 (A), tnni1a (C), mylz10 (E), tpm2 (G) and ryr1a (I) is downregulated in slow muscle fibers ectopically expressing Sox6-GFP (B,D,F,H,J). |
|
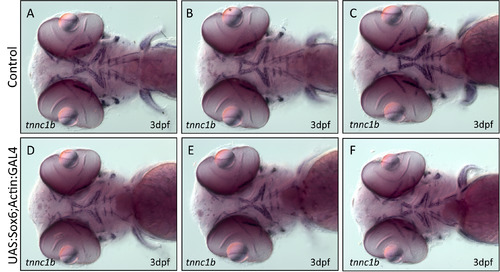
Misexpression of Sox6 has no effect on the expression of tnnc1b in the head muscles. (A-C ) tnnc1b expression in the head muscles of 3 dpf wild type embryos. (D-F) tnnc1b expression in embryos expressing Sox6-GFP in head muscles. There appears to be no difference in the expression of tnnc1b between wild type and actin:GAL4:UAS:sox6-GFP fish. |
|
GFP expression is unaffected by mutation of potential Sox6 binding sites in the +2.5kb + intron1 tnnc1b:eGFP reporter. Expression remains restricted to the slow-twitch domain following mutation of one or more putative Sox6 binding sites (A-F). |
|
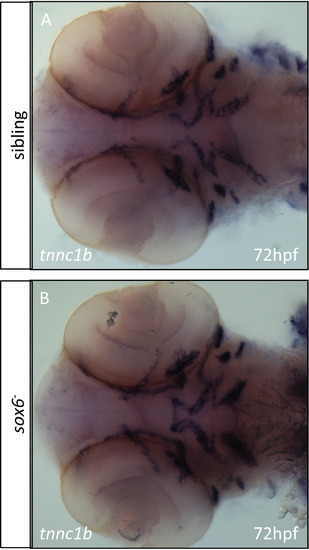
In situ hybridization for tnnc1b mRNA reveals no difference in expression between wild type and sox6 mutants in the craniofacial muscles. |
|
Slow-specific muscle genes show a variable response to mutation of sox6. Like tnnc1b and ryr1a, tnnt1 is ectopically expressed throughout the fast domain of sox6 homozygous mutant embryos at 30 hpf (A and F). Conversely prox1a expression is unaffected in sox6 mutant embryos at 22 hpf when transcript levels can be detected (B and G). Expression of tnni1a (C and H), mylz10 (D and I) and tpm2 (E and J) is unaffected at 30 hpf in sox6 mutants. |
|
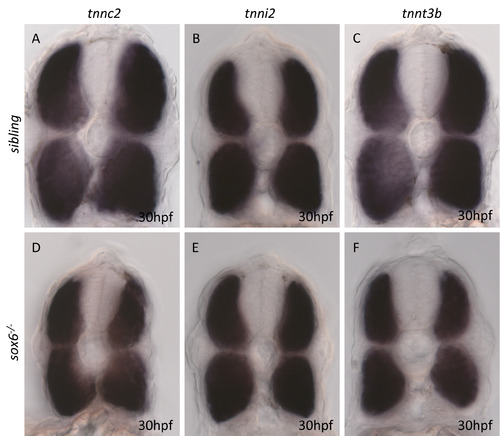
Expression of fast specific troponin subunit genes in wild type and sox6 mutant embryos. |
|
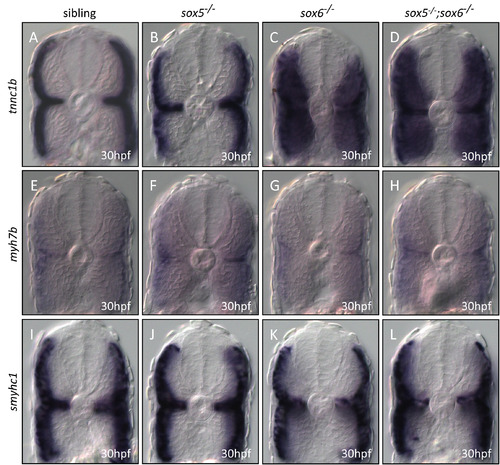
Expression of smyhc1, myh7b and tnnc1b in wild type, sox5, sox6 and sox5; sox6 double mutants. |