- Title
-
Polarization and orientation of retinal ganglion cells in vivo
- Authors
- Zolessi, F.R., Poggi, L., Wilkinson, C.J., Chien, C.B., and Harris, W.A.
- Source
- Full text @ Neural Dev.
|
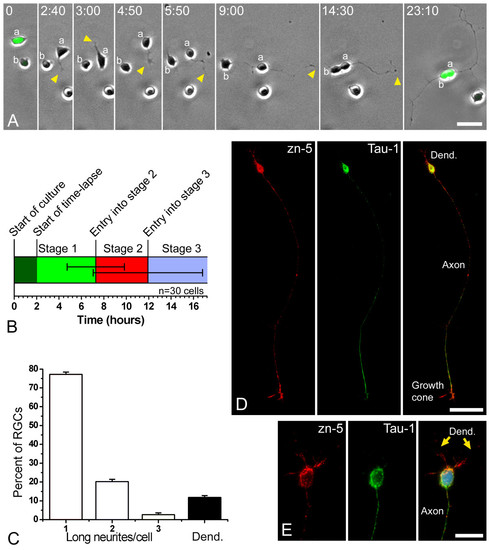
Retinal ganglion cells (RGCs) polarize in vitro after a period of plastic behavior. (a) Time-lapse analysis of dissociated ath5:gfp-expressing retinal cells in culture (Additional file 1). At the start of the time-lapse, cell 'a' expressed a higher level of GFP and was already in late stage 1 of differentiation (namely forming long filopodia), whereas cell 'b' showed a lower fluorescence and was at early stage 1. A short time later cell 'a' started to generate short processes ('neurites'; arrowheads), indicating the onset of stage 2. One of the neurites formed a growth cone at time point 4 minutes 30 seconds and started to grow faster at the beginning of stage 3. After the cell bodies made contact, near the end of the sequence, cell 'b' also seemed to extend an axon-like neurite. Time is shown in hours:minutes. Scale bar, 15 μm. (b) Graphic representation of the timing of in vitro differentiation of zebrafish RGCs, where 30 cells were followed by time-lapse video microscopy. The horizontal lines in the middle of the bar represent the standard deviation from the transitions between stages 1 and 2 and between stages 2 and 3. (c) Analysis of the morphology of the RGCs (Zn5-positive cells) after 24 hours in culture. n = 100 cells, in three independent cell-culture experiments. 'Long neurites' are longer than three cell diameters. (d, e) Cultured RGCs at stage 4 (24 hours in vitro), labeled with the RGC-specific antibody Zn-5 (red), an anti-Tau-1 antibody (green) and 4',6-diamidino-2-phenylindole in (e). Scale bars, 30 μm (d) and 10 μm (e). |
|
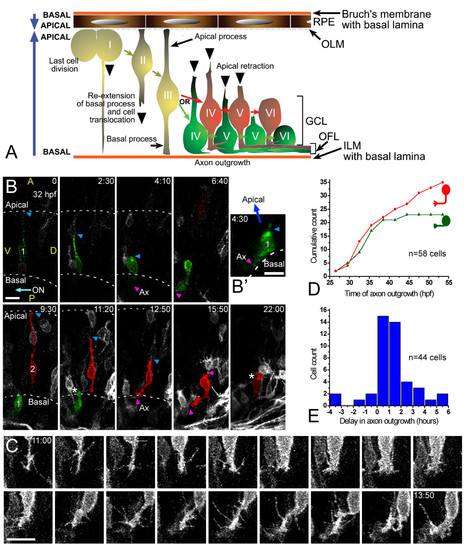
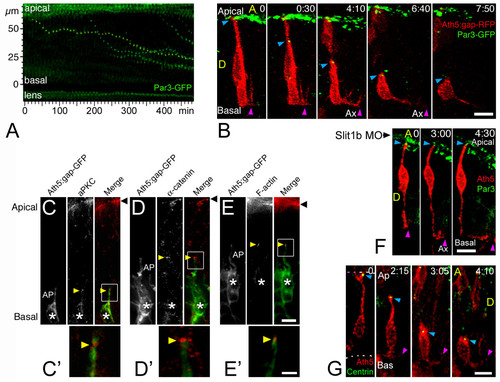
Retinal ganglion cell (RGC) polarization in vivo seems restricted by the environment. (a) Summary of the observed behaviors of normal RGCs in vivo. GCL, ganglion cell layer; ILM, inner limiting membrane; OFL, optic fiber layer; OLM, outer limiting membrane. (b) Time-lapse confocal (four-dimensional (4D)) analysis of an ath5:gap-gfp-injected embryo, showing two differentiating cells that undergo apical retraction and axonogenesis (Additional file 2). Cell 1 (green) shows no basal process when forming the axon (see view rotated 90° on the y axis in (b')), whereas cell 2 (red) does. Ax, axon; ON, optic nerve; A, anterior, D, dorsal; P, posterior; V, ventral; the asterisk marks cell processes at the apical side of the differentiating RGCs, where the dendrites are forming. The recording was started at 32 hpf; time is shown in hours:minutes. Scale bars, 10 μm. (c) Complete sequence of images from cell 2, with images taken every 10 minutes and showing a detail of the basal cell surface dynamics just before axonogenesis. Scale bar, 10 μm. (d) Cumulative plot showing the time of axon formation for RGCs with (red dots) and without (green triangles) a visible basal process at the moment of forming the axon, from the 4D analyses in vivo. (e) Distribution analysis of the delay between apical retraction and axonogenesis in 44 cells followed by 4D imaging. |
|
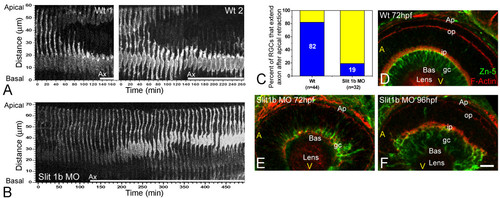
The timing of axon formation does not correlate with the dynamics of apical retraction. (a) Time-series images of the cells shown in Figure 2b, showing the retraction of the apical process in a wild-type embryo (also see Figure 3). (b) A similar analysis performed in an ath5:gap-gfp transgenic embryo, injected with 2 ng of morpholino to Slit1b. The cell is able to extend an axon (not shown in the picture; see Additional file 3) but does not retract its apical process during the time of the movie (more than 8 hours). The age of the embryo at the start of the movie was about 48 hpf. The white bars under the time-series images indicate the presence of an axon (Ax, not seen in the pictures). (c) Comparison of the percentage of retinal ganglion cells (RGCs) forming their axons before or after the detachment of the apical process in wild-type and Slit1b morpholino-injected embryos, as analyzed by four-dimensional microscopy; n is the number of cells followed in 12 wild-type and 7 morphant embryos. (d-f) Parasagittal optical sections of wild-type and Slit1b-treated embryos, labeled for RGCs (gc) with Zn-5, and for F-actin with phalloidin-Texas red, which predominantly stains the inner plexiform (ip) and outer plexiform (op) layers, and the outer limiting membrane (Ap). Bas, basal; A, anterior, V, ventral. Scale bar, 10 μm. EXPRESSION / LABELING:
|
|
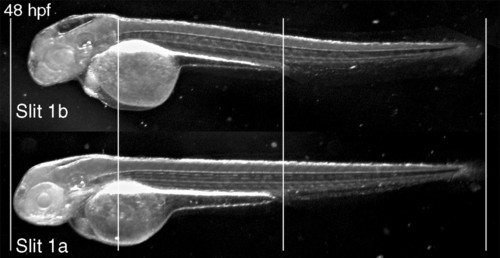
General phenotype of Slit1b morphants. External morphology of a Slit1b morphant at 48 hpf, compared with an embryo treated with the same amount (2 ng) of Slit1a morpholino, which is indistinguishable from a wild-type embryo (not shown). The alignment of the embryos on the vertical lines shows how the general growth of the Slit1b morphants is not affected. The head region, in contrast, seems severely affected, being smaller overall and having smaller eyes and thinner brain walls (with bigger ventricles). PHENOTYPE:
|
|
The apical process of retinal ganglion cells (RGCs) remains with apical identity during axon initiation. (a) Time-series representation of the Par3-GFP-labeled tips of retracting apical processes. Two dots (from contiguous cells) that show a complete movement from their initial (apical) to their final (basal) positions are pseudo-colored in yellow and blue (Additional file 4). The movement of these dots is reminiscent of that of the retraction of the apical processes (see Figure 3). Time point 0 is at 32 hpf. (b) Four-dimensional (4D) analysis of an embryo expressing ath5:gap-rfp (injected as plasmid DNA) and Par3-GFP (injected as mRNA). The blue arrowhead indicates the tip of the apical process of a differentiating RGC undergoing axonogenesis, and the purple arrowhead indicates the axon (Additional file 5). The green signal seen at the top of the images represents the Par3-GFP accumulation at the apical border of retinal neuroepithelial cells and of retinal pigment epithelium cells. Time point 0 is at 32 hpf. Ax, axon; A, anterior, D, dorsal; time is shown in hours:minutes. Scale bar, 10 μm. (c-e) Immunolabeling of ath5:gap-gfp transgenic embryos with different apical markers (anti-aPKC and anti-α-catenin antibodies, and phalloidin-Texas Red to label actin filaments), showing their accumulation (yellow arrowheads) at the tip of retracting apical processes (AP) of RGCs (asterisks). Eight to ten embryos, at different stages, were used in this analysis for each marker, and representative examples are shown. (c,d) 33 hpf; (e) 42 hpf. Black arrowhead: apical border of the neuroepithelium. Scale bars, 5 μm (c-e) and 2 μm (c'-e'). (f) 4D sequence from an ath5:gap-rfp (transgenic) and Par3-GFP (mRNA-injected)-expressing embryo, treated with Slit1b morpholino. Note the formation and elongation of the axon (purple arrowhead) without detachment of the apical process from the apical side of the neuroepithelium (blue arrowhead). Time point 0 is at 40 hpf. Ax, axon; time is shown in hours:minutes. Scale bar, 10 μm. (g) 4D analysis of an embryo expressing ath5:gap-rfp (injected as plasmid DNA) and GFP-zcentrin (injected as mRNA). The blue arrowhead indicates the tip of the apical process of a differentiating RGC, and the purple arrowhead indicates the axon (Additional file 6). Some green background appears in the pictures, which mostly comes from cytoplasmic GFP-zcentrin in cells that express a very high level of the fusion protein (this is not the case in the highlighted cell). That the signal is predominantly centrosomal can be seen better in Additional file 6. Time point 0 is at 32 hpf. Ax, axon; A, anterior, D, dorsal; time is shown in hours:minutes. Scale bar, 8 μm. |
|
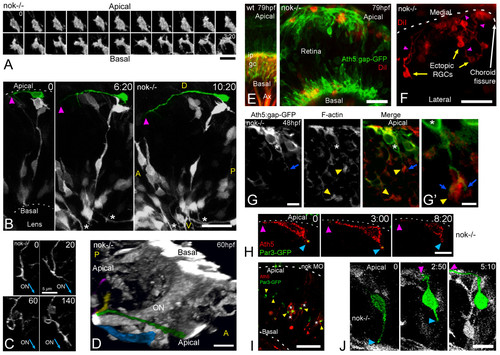
Retinal ganglion cells (RGCs) polarize, but can be inverted in nok mutants. (a) Complete sequence of images from a three-dimensional (3D) reconstruction, seen from a lateral-apical perspective, of an ectopic RGC in a nok-/- retina. Just before axon outgrowth, the RGC presents several filopodia, mostly extending from the site of axon formation (compare with Figure 2c). Scale bar, 5 μm. (b) Four-dimensional (4D) analysis of a nok-/-embryo injected with ath5:gap-gfp DNA. The ectopic RGC pseudo-colored in green is extending a long neurite (arrowhead) on the retinal outer surface. Time point 0 is at 32 hpf. Asterisks indicate axons from basally located RGCs; A, anterior, D, dorsal; P, posterior; V, ventral. Scale bar, 25 μm. (c) Sequence of 3D reconstructions from a nok-/-embryo expressing ath5:gap-gfp, where an axon growing on the retinal outer surface loops before continuing its growth apparently directed towards the optic nerve (ON) exit. Scale bar, 5 μm. (d) 3D reconstruction taken from a 4D analysis of a nok-/- embryo, transgenic for ath5:gap-gfp. The eye is seen from a medio-ventral position. The pseudo-color highlights four ectopic fascicles of axons joining the optic nerve outside the retina (Additional file 7). Scale bar, 10 μm. (e) Extended-focus confocal images of ath5:gap-gfp transgenic embryos injected with the lipophilic dye DiI into the right tectum to label RGCs retrogradely in the left eye. Scale bar, 15 μm. (f) Ventral view of an eye from a nok-/- embryo in which RGCs have been retrogradely labeled with DiI. The axons of the ectopic RGCs are seen growing towards the optic nerve (not shown in the picture) on the outer retinal surface. Scale bar, 25 μm. (g) Optical section of an ath5:gap-gfp transgenic, nok-/- retina, stained with phalloidin-Texas red. The 'apical' process (blue arrow) of an ectopic ath5:gap-gfp cell (asterisk) ends close to an area where actin filaments appear accumulated (yellow arrowhead). (g' ) Higher magnification of the same cell. Scale bars, 5 μm (g) and 2 μm (g'). (h) Sequence of optical sections from a 4D analysis of a nok-/- retina expressing ath5:gap-rfp and Par3-GFP. The ath5:gap-rfp-positive cell is located close to the apical surface of the retina and is extending a neurite on the outer retinal surface (purple arrowhead). On its other end, another process shows an accumulation of Par3-GFP in its tip (blue arrowhead). Time point 0 is at 34 hpf. Scale bar, 10 μm. (i) Optical section of a retina from a living nok-/- embryo injected with ath5:gap-rfp DNA and par3-gfp mRNA. The Par3-GFP protein is accumulated in apparently random positions inside the retina, and the ath5:gap-rfp-positive cells can be found either on the apical or basal sides of these accumulation points. Scale bar, 20 μm. (j) Sequence of extended-focus images from a 4D analysis of a nok-/-, ath5:gap-gfp transgenic embryo, in which an ectopic differentiating RGC (highlighted in green) is seen to retract a basally directed process and then extend an axon on the retinal outer surface (see Additional file 8 for a rotated 3D reconstruction of this cell). Time point 0 is at 32 hpf. Scale bar, 10 μm. All time stamps are in hours:minutes. |
|
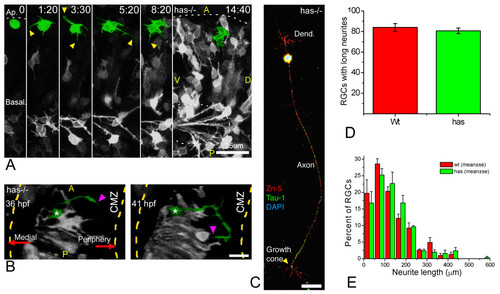
Polarization of ectopic RGCs is impaired in a non cell-autonomous manner in has mutants. (a) Four-dimensional (4D) analysis of the retina of a has-/- embryo injected with ath5:gap-gfp plasmid DNA (Additional file 9). The cell highlighted in green divides at the first time frame, and the two daughter cells remain for several hours at the apical region of the retina, increasing their GFP expression but not generating any long neurites. Time point 0 is at 32 hpf. A, anterior, D, dorsal; P, posterior; V, ventral. Scale bar, 25 μm. (b) A dorsal view of a three-dimensional reconstruction taken from a 4D confocal analysis of a has-/- embryo transgenic for ath5:gap-gfp. The ectopic retinal ganglion cell (RGC) highlighted in green is growing a long neurite (arrowheads) that is initially directed towards the retinal periphery (that is, opposite to the optic nerve exit), and then turns back towards the cell. CMZ, ciliary marginal zone. Scale bar, 15 μm. (c) has-/- RGC in culture, triple-labeled with Zn-5 and anti-Tau-1 antibodies and 4',6-diamidino-2-phenylindole. The cell is indistinguishable from a stage 4 wild-type RGC (see Figure 1). Scale bar, 15 μm. (d,e) Quantitative analysis of neuritic outgrowth in vitro from has-/-, compared with wild-type, RGCs (defined as Zn-5-positive cells). (d) Number of RGCs with long (more than three cell diameters) neurites; n = 200 cells per strain from three independent experiments. (e) Distribution of neurite lengths; n = 100 neurites per strain (one neurite per cell; the longest was chosen in cells with more than one) from three independent experiments (normalized values). |
|
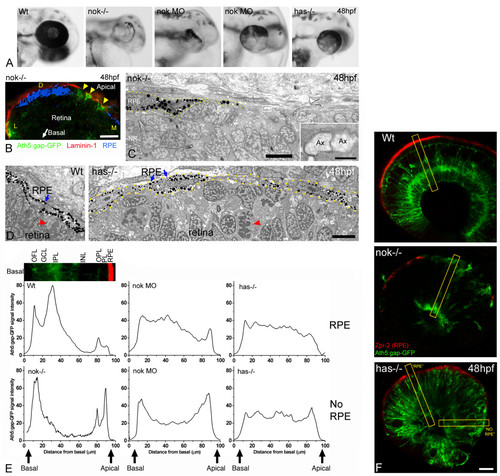
The RPE influences the ability of retinal ganglion cells to polarize in ectopic positions. (a) Head region of wild-type, morphant and mutant embryos, showing the distribution of pigment around the retina. Two examples of nok morpholino-injected embryos (0.17 pmoles per embryo) are shown. (b) Cryosection of the eye from a nok-/-, ath5:gap-gfp transgenic embryo, labeled with an anti-laminin 1 antibody (red). The bright-field image, inverted and pseudo-colored in blue, shows the distribution of the retinal pigment epithelium (RPE). Arrowheads indicate ath5:gap-gfp-positive retinal ganglion cells. D, dorsal; L, lateral; M, medial. Scale bar, 20 μm. (c) Transmission electron micrograph of the apical region of a nok mutant retina, showing the distribution of the RPE cells (dotted lines). Inset, a higher magnification of the boxed region, showing two apparent transverse-sectioned axons (Ax) at the RPE-free apical surface of the retina. Scale bars: low-magnification image, 5 μm; inset, 0.5 μm. (d) Transmission electron micrographs of the apical region of a wild-type (Wt) and a has-/- embryo. The RPE (blue arrows) is a very organized simple epithelium in the wild type, and it seems disrupted in the has mutant. Nevertheless, the picture does not show any actual gap between the RPE cells in the has-/-embryo. Red arrowheads indicate mitotic cells in a normal (apical) position in the wild-type, and in an ectopic position in the mutant. Scale bar, 5 μm. (e) Fluorescence intensity profiles made on confocal sections of 48 hpf ath5:gap-gfp transgenic embryos labeled with an anti-RPE antibody (Zpr-2). For the measurements a 20-pixel wide line was drawn along a radius of the retina as in (f). The intensity profile of the green channel (GFP) was plotted and the values were normalized (maximum intensity) and averaged for each embryo; the resulting plots were then normalized to each other (integrated intensity). To compare the profiles in relation to the distribution of the RPE, we positioned the line either in regions where zpr-2 immunoreactivity was detected (RPE) or not (NO RPE). For the nok mutants, only measurements of areas without detectable RPE were used. Measurements were made in three to ten different areas from one wild-type, five nok mutant, four nok morphant and three has mutant retinas. The inset picture shows a region of a wild-type retina, like those used for the measurements, which is aligned with the profile plot to show the correspondence with the retinal layers. GCL, ganglion cell layer; INL, inner nuclear layer; IPL, inner plexiform layer; OFL, optic fiber layer; OPL, outer plexiform layer; PL, photoreceptor layer. (f) Optical sections of wild-type and mutant retinas of ath5:gap-gfp transgenic embryos labeled in red with the Zpr-2 antibody, which stains the RPE. The yellow rectangles show an example of the lines used for the measurements presented in (e). Scale bar, 25 μm. |
|
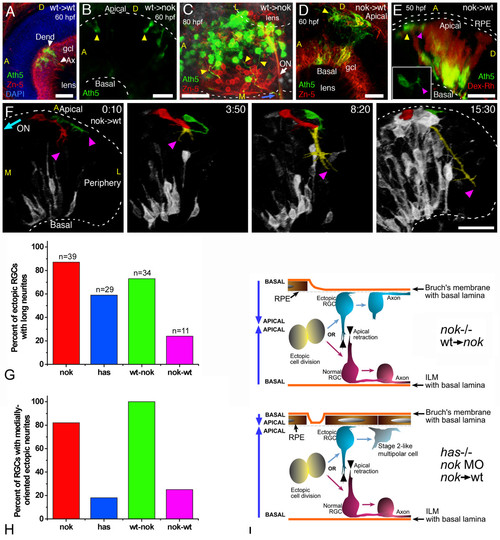
Importance of the tissue environment for the differentiation and orientation of ectopic retinal ganglion cells (RGCs). (a) Extended-focus confocal image from a wild-type host embryo transplanted with blastomeres from an ath5:gap-gfp transgenic wild-type embryo. gcl, ganglion cell layer. (b,c) Extended-focus images from nok-/- host retinas transplanted with ath5:gap-gfp transgenic cells (wild type). In (b), the lateral view of the eye shows the presence of ectopic ath5:gap-gfp-positive cells (arrowheads); in (c), the eye is seen from a ventral view, and many ectopic (apical) donor RGCs that extend axons (yellow arrowheads) joining the optic nerve (ON) on the outside of the eye are shown. (d-f) Images of wild-type hosts transplanted with blastomeres from nok-/-, ath5:gap-gfp transgenic donors. (d) Extended-focus confocal image, in which the donor RGCs (positive for both Zn-5 (red) and ath5:gap-gfp (green)) are found either mixed with host RGCs in a normal-appearing ganglion cell layer (gcl) or on the apical side of the retina. (e) Rotated extended-focus confocal image, showing how a nok-/- donor-derived ectopic (apical; yellow arrowhead) ath5:gap-gfp-positive cell starts to extend a neurite (purple arrowhead) that will grow only a few micrometers before turning back to the originating cell (Additional file 10). The red stain shows all the donor cells, labeled with dextran-tetramethylrhodamine β-isothiocyanate (Dex-Rh). No transplanted cells are detected in the surrounding retinal pigment epithelium, indicating that it must all be of wild-type (host) origin. The inset shows a better view (from the apical side, in this case in an optical section orthogonal to the confocal laser) of the cell in the same time point. (f) Sequence of rotated three-dimensional reconstructions taken from a four-dimensional analysis. The two cells highlighted are from the donor (nok-/-) and are extending long neurites, which will grow together towards the retinal periphery (colored yellow where they are indistinguishable from each other). Time point 0 is at 32 hpf. ON, optic nerve; A, anterior; D, dorsal; L, lateral; M, medial. All scale bars are 25 μm long. (g) Comparison of the number of apically located ath5:gap-gfp cells extending a long neurite (longer than three cell diameters), in different mutant and morphant conditions. Total number of cells (n) and embryos analyzed (cells/embryos): nok-/-, 39/11; has-/-, 29/8; transgenic nok-wild-type, 34/9; transplanted wild-type-nok, 11/6. (h) The proportion of these cells in which the neurite is growing towards the medial region of the eye, where they would be able to meet the optic nerve. (i) Summary of the observed phenotypes of ectopic RGCs in mutant, morphant and transplantation conditions. |