- Title
-
Pcdh18a regulates endocytosis of E-cadherin during axial mesoderm development in zebrafish
- Authors
- Bosze, B., Ono, Y., Mattes, B., Sinner, C., Gourain, V., Thumberger, T., Tlili, S., Wittbrodt, J., Saunders, T.E., Strähle, U., Schug, A., Scholpp, S.
- Source
- Full text @ Histochem. Cell Biol.
|
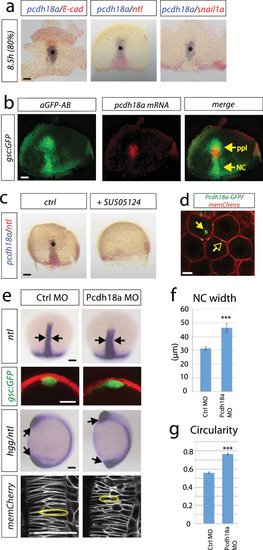
A cell group in the axial mesoderm regulates notochord morphogenesis in zebrafish. EXPRESSION / LABELING:
PHENOTYPE:
|
|
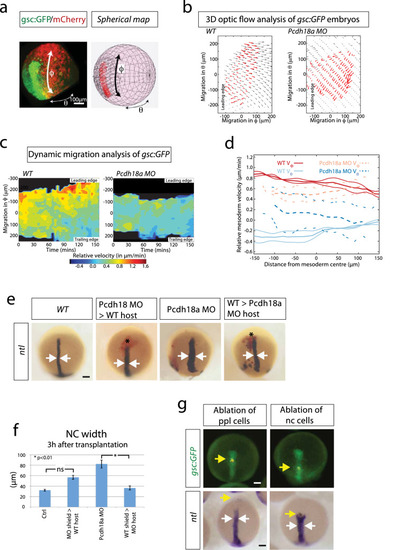
Analysis of the influence of the ppl on notochord morphogenesis. EXPRESSION / LABELING:
PHENOTYPE:
|
|
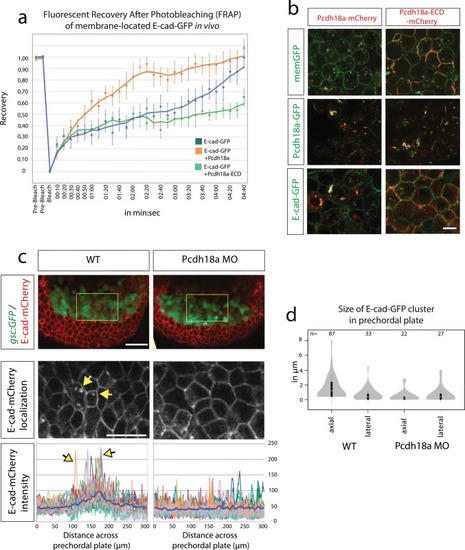
Pcdh18a regulates recycling of the E-cadherin. |
|
Pcdh18a domains and their importance with regard to endocytosis. |
|
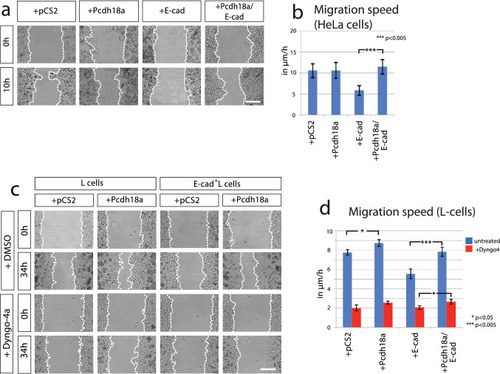
Pcdh18a affects E-cadherin dependent cell migration. |
|
Directed cohort migration results in the formation of the rod-shaped notochord. |