- Title
-
Regulatory Network of the Scoliosis-Associated Genes Establishes Rostrocaudal Patterning of Somites in Zebrafish
- Authors
- Keskin, S., Simsek, M.F., Vu, H.T., Yang, C., Devoto, S.H., Ay, A., Özbudak, E.M.
- Source
- Full text @ iScience
|
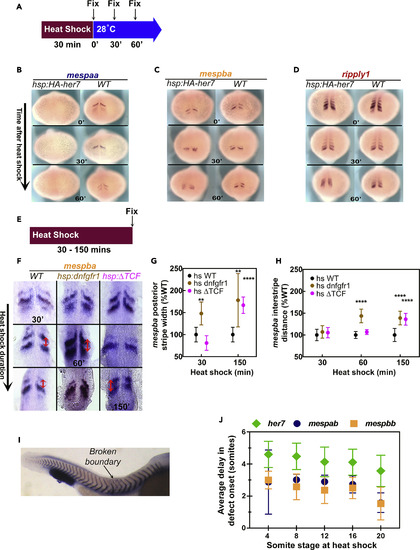
Expression of mesp Genes Read Out the Segmentation Clock Genes and FGF Signaling (A) Embryos from different genetic backgrounds were fixed immediately or after 30 min, or 60 min of recovery following 30-min heat shock at 37°C. (B–D) ISH images of mespaa (B), mespba (C), and ripply1 (D) after heat shock of tg(hsp70l:HA-her7) and wild-type (WT) embryos at different recovery time points. This experiment was repeated twice and 43–49 embryos analyzed for all probes and time points. (E) Embryos from different genetic backgrounds were fixed immediately after 30, 60, 90, 120, or 150 min of heat shock at 37°C. (F) Flat mounted ISH images of mespba after heat shock of tg(hsp70l:dnfgfr1a-EGFP), tg(hsp70l:tcf7l1a-GFP), and wild-type (WT) embryos at different time points. For each time point 8 to 22 embryos were analyzed. Red arrows show the interstripe distance, which was measured between the anterior ends of stripes. (G and H) Effect of inhibition of FGF (brown) or Wnt (pink) signaling on the width of the posteriormost stripe of mespba expression stripe (G) and the interstripe distance between two stripes of mespba expression (H) shown as percent of that in wild-type embryos exposed to heat shock. Error bars indicate one standard deviation. (I) ISH image of xirp2 showing position of the somiteboundary defects in a tg(hsp70l:HA-her7) embryo after 40 min of heat shock. Heat shock started when embryo was at 4-somite stage; first broken boundary appears between 9th and 10th somites as indicated. (J) Average delay in onset of segmentation defects in embryos overexpressing her7 [tg(hsp70l:HA-her7)], mespab[tg(hsp70l:mespab-myc)], or mespbb [tg(hsp70l:mespbb-myc)] at different somite stages after heat shock at 37°C. her7transgenic embryos were subjected to heat shock for 40 min; mespab and mespbb transgenic embryos were subjected to heat shock for 30 min. Experiments were repeated twice, and 30–62 embryos were analyzed for all genotypes and stages. Error bars indicate one standard deviation. |
|
Gene Regulatory Network in the Anterior PSM Is Mapped (A) Wild-type (WT) embryos were continuously treated in DAPT or DMSO and fixed at different time points. (B–D) ISH pictures for (B) mespaa, (C) mespba, and (D) her7genes in DAPT- or DMSO-treated WT embryos. These experiments were repeated twice, and 38–52 embryos were analyzed for all conditions, probes, and stages. (E) Embryos from different genetic backgrounds were fixed immediately after 60 min of heat shock at 37°C. (F and G) ISH images of the expression pattern of her7and deltaC immediately after 1-h heat shock in (F) hsp70l:mespab-myc or (G) hsp70l:mespbb-myc embryos. (H and I) ISH images of mespba or mespaa expression immediately after 1-h heat-shock in (H) hsp70l:mespab-myc or (I) hsp70l:mespbb-myc embryos. (F–I) These experiments were repeated twice, and 50–65 embryos were evaluated for all genotypes, probes, and stages. |
|
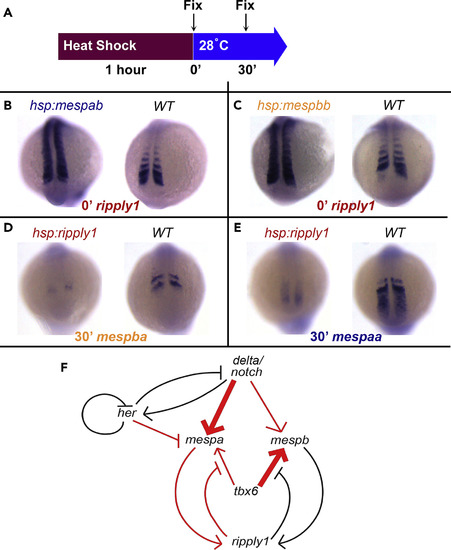
There Is a Negative Feedback Loop between mespa/mespb and ripply1 Genes (A) Embryos from different genetic backgrounds were fixed immediately or 30 min of recovery after 60-min heat shock at 37°C. (B and C) ISH hybridization pictures of ripply1 gene in (B) hsp70l:mespab-myc or (C) hsp70l:mespbb-myc embryos after a 1-h heat shock. These experiments were repeated three times, and 104–96 embryos were evaluated for each genotype, respectively. (D and E) ISH hybridization pictures of (D) mespba and (E) mespaa in hsp70l:ripply1-myc embryos after a 1-h heat shock. These experiments were repeated twice and 45–47 embryos were evaluated for all probes. (F) Regulatory network among scoliosis-linked genes. herrepresents both her1 and her7. delta/notch represents the transcriptional activation of its ligand DeltaC and subsequent pathway activation. Thicker arrows indicate stronger and differential dependence of transcription of each mesp gene on two different transcription factors. Red color represents regulatory interactions inferred from time-controlled perturbation experiments in this study. In the computational model, transcription, translation, protein and mRNA degradation, formation of protein complexes, and proteolytic Notch activation are included for the nodes in the network. WT, wild-type. |