- Title
-
Concerted action of neuroepithelial basal shrinkage and active epithelial migration ensures efficient optic cup morphogenesis
- Authors
- Sidhaye, J., Norden, C.
- Source
- Full text @ Elife
|
RNE invagination is accompanied by basal cell surface area shrinkage and basal actomyosin accumulation. (A) Schematic representation of RNE morphogenesis from 16 hours post fertilization (hpf) or 15 somite stage (ss) to 24 hpf or 30 ss showing RNE cells that undergo invagination (brown) and rim cells that undergo rim involution (blue). Dotted arrow marks direction of rim involution. (B) Average area of RNE cells at apical (magenta) and basal (green) sides during RNE morphogenesis with mean ± SD. N = 10 embryos. See Figure 1—source data 1. (C,D) Time-lapse imaging of RNE morphogenesis to assess the dynamics of actin marked by GFP-UtrophinCH (C) and of myosin marked by myl12.1-EGFP (D). Arrowhead marks basal actin and myosin enrichment in the RNE. Arrow marks the rim zone lacking the basal enrichment. Videos started around 15 ss. Time in h:min. Frames from Videos 1 and 2. (E,G) Confocal scan of optic cup at 30 ss immunostained for phalloidin (E) and phosphomyosin (G). The arrowhead marks enrichment at the basal side of the RNE. Lookup table indicates maximum and minimum intensity values. (F,H) Normalized average intensity distributions of phalloidin (F) and phosphomyosin (H) in the tissue volume along the apicobasal axis of the RNE at 30 ss. Mean ± SEM. The schematic shows a typical tissue section used for analysis (Also see Figure 1—figure supplement 1D). Tissue sections, n = 14 for phalloidin and n = 15 for phosphomyosin; N = 5 embryos. See Figure 1—source data 2 and 3. All scale bars = 10 µm. Developmental axes are shown next to the panels for orientation. D: Dorsal, V: Ventral; Prox: Proximal, Dist: Distal; A/N: Anterior or nasal, P/T: Posterior or temporal. |
|
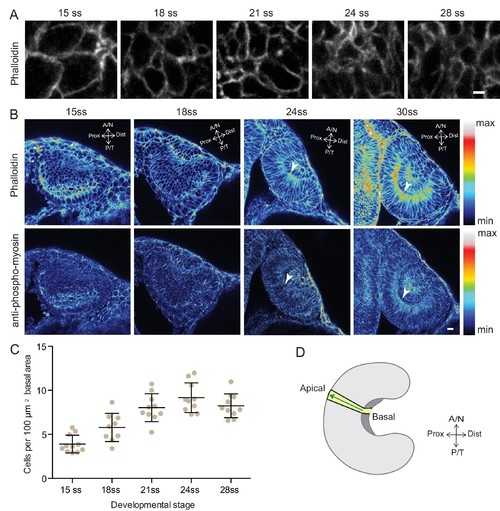
Analysis of the basal surface and actomyosin localisation during RNE invagination. (A) Confocal scans of the basal side of the RNE cells stained with phalloidin at different developmental stages.Scale bar = 2 µm. (B) Confocal scans of different stages of RNE morphogenesis stained for phalloidin and phosphomyosin. Arrowheads point at basal enrichment. Lookup table indicates the minimum and maximum intensity values. Scale bar = 10 µm. (C) Average number of cells per 100 µm2 basal area of RNE at different stages of development. Mean ± SD, N = 10 embryos each. See Figure 1—source data 1. (D) Schematic drawing of a typical tissue volume used for actomyosin intensity distribution analysis (yellow). The average intensity was measured from basal to apical side of the tissue volume (green arrow) and normalised to the highest intensity value along the axis. See Materials and methods for details. |
|
Impairment of actomyosin-driven basal constriction or proliferation does not prevent RNE formation. (A) Confocal scan of optic cup at 30 ss stained for phalloidin. Control (left), phenotypes after 7 h of Rockout treatment: invagination defect in 30% embryos (middle), invagination defect with epithelial accumulation in 40% embryos (right), (n = approx. 30 embryos, N = 5 experiments, see Figure 2—source data 1). Arrowheads mark the basal side of the RNE. Arrow marks the epithelial accumulation outside the RNE. Asterisk marks the developing lens. Dashed lines indicate the angle of invagination. Rockout treatment started around 13–14 ss. Lookup table indicates the minimum and maximum intensity values. Scale bar = 10 µm. (B) Brightfield images of optic cup at the end of the 7 h treatment. Control (left), treated with Rockout (right). Arrow marks the epithelial accumulation outside the RNE. Scale bar = 50 µm. (C,D) Normalized average intensity distributions of phalloidin (C) and phosphomyosin (D) in the tissue volume along the apicobasal axis of the RNE at 30 ss. Mean ± SEM. Control (brown), Rockout treated (grey). Tissue sections, n = 25; N = 5 embryos each. See >Figure 2—source data 2 and 3. (E) Time-lapse of RNE morphogenesis in DMSO Control (upper) and Rockout-treated (lower) embryos expressing actin marker GFP-UtrophinCH. N = 5 embryos each. Rockout treatment was started 2 h before imaging around 13–14 ss. Imaging started at around 18 ss. Both movies were imaged simultaneously and under identical imaging conditions. Time in h:min. Frames from Video 3. Scale bar = 10 µm. (F) Invagination angle at 30 ss. Mean ± SD. The schematic shows the invagination angle as the angle held at the base of central RNE by the inner lips of the optic cup. P values for Mann-Whitney test with Control are as follows: Rockout p=0.0004, HU+Aphi p=0.0421 and for laminin a1 Mo, opo Mo and ezrin Mo p<0.0001. See Figure 2—source data 4. (G) Average basal area of RNE cells with mean ± SD. Each dot represents one embryo. P values for Mann-Whitney test with 30 ss control are as follows: 30 ss Rockout p=0.0952, 30 ss HU+Aphi p<0.0001. See Figure 2—source data 5. |
|
Effect of Rockout and HU+Aphi treatment on the RNE. (A) Confocal scan of 30 ss RNE in control (left) and Rockout treated (right) embryo stained for phosphomyosin.Rockout treatment was performed for 7 h starting from 13–14 ss. Lookup table indicates the minimum and maximum intensity values. (B) Confocal scans of 30 ss RNE in control (left) and HU+ Aphi treated (right) embryos stained for DAPI (cyan), mitotic marker phosphohistone-3 (yellow) and phalloidin (magenta). (C) Confocal scans of 30 ss RNE in control (left) and HU+ Aphi-treated (middle and right) embryos stained with phalloidin. Areas marked by yellow box are shown in D. (D) Confocal scans of RNE cells (zoomed images of regions marked in C). Control (top), HU+Aphi treated (middle and bottom) stained with phalloidin. All scale bars = 10 µm. |
|
Rim involution involves active cell migration of connected epithelial cells. (A) Time-lapse imaging of rim zone with mosaic expression of Par3-GFP and mKate2-ras. Arrows show membrane protrusions. Frames from Video 4. N = 5. (B) Time-lapse imaging of rim zone with mosaic expression of GFP-UtrophinCH and ras-mKate2. Inlays show zoomed marked area. Arrows show actin localization in the protrusions. Arrowheads mark the basally enriched stable actin pool in the RNE. Frames from Video 5. N = 6. (C) Time-lapse imaging of rim zone with mosaic expression of GFP-UtrophinCH and ras-mKate2. Yellow arrows show apical actin localization at adherens junctions. Frames from Video 7. N = 6. (D) Schematic of a rim cell exhibiting actin based protrusions at the basal side. The arrow marks the direction of involution. The leading and lagging edges indicate the sides referred in (E) and (F). (E) Number of actin protrusions observed per hour in rim cells in the control condition. Each pair of datapoints represents two sides of the same rim cell. n = 9, N = 6 . See Figure 3—source data 1. (F) Number of actin protrusions observed per hour in the rim cells in Rockout treatment condition. Each pair of datapoints represents two sides of the same rim cell. n = 6, N = 6. See Figure 3—source data 1. (G) Confocal scan of rim zone in Rockout-treated embryos with mosaic expression of GFP-UtrophinCH and ras-mKate2. Inlays show enlarged marked area. Arrows show actin localization in the protrusions. Frames from Video 6. N = 6 all scale bars = 10 µm. All movies started around 17 ss -18 ss, Time in h:min. |
|
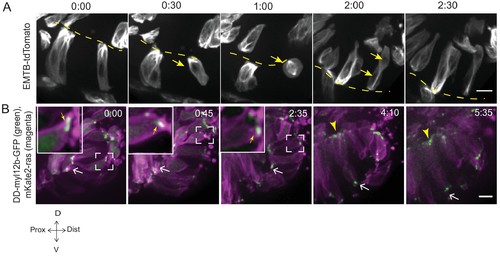
Microtubule and myosin dynamics during rim involution. (A) Time-lapse imaging of rim zone with mosaic expression of EMTB-tdTomato.Arrows mark the cell dividing during rim involution. Dashed line marks the apical side. Imaging started at 17–18 ss. N = 4. (B) Time-lapse imaging of rim zone with mosaic expression of DD-myl12b-GFP and mKate2-ras. Inlays show zoomed marked area. Yellow arrows indicate basal myosin punctae in the rim cells. White arrows mark adherens junctions. Arrowheads mark basally enriched stable myosin pool in the RNE. Frames from Video 9. Imaging started at 15–16 ss. N = 5. Scale bars = 10 µm, Time h:min. |
|
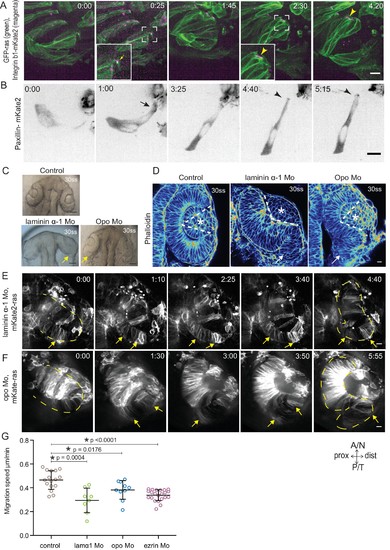
Dynamic cell-ECM attachment of rim cells is important for RNE morphogenesis. (A) Time-lapse imaging of rim zone with mosaic expression of GFP-ras and Integrin-β1b-mKate2. Inlays show enlarged marked area. Arrow indicates the integrin foci during migration. Arrowheads mark the basally enriched stable integrin pool in the RNE cell. Frames from Video 10. N = 4. Scale bar = 10 µm. Imaging started at 17–18 ss, Time in h:min. (B) Timelapse imaging of rim zone with mosaic expression of paxillin-mKate2. Arrow indicates the short-lived paxillin foci. Arrowhead marks the stable basal paxillin localization in the RNE. Frames from Video 11. N = 6. Scale bar = 10 µm. Imaging started at 17–18 ss. Time in h:min. (C) Brightfield images of the dorsal view of 30 ss embryo head. Control (upper), laminin morphant (lower left) and opo morphant (lower right). Arrows mark the epithelial accumulation outside the RNE. Scale bar = 50 µm. (D) Confocal scan of optic cups at 30 ss stained for phalloidin. Control (left), laminin morphant (middle) and opo morphant (right). Arrows mark the epithelial accumulation outside the RNE. Dashed lines indicate the angle of invagination. Asterisk marks the developing lens. Lookup table indicates the minimum and maximum intensity values. Scale bar = 10 µm. (E,F) Time-lapse imaging of RNE morphogenesis in laminin morphant (E) and Opo morphant (F) injected mosaically with mKate2-ras RNA. Arrows mark rim cells that failed to move. Dashed line marks the outline of the developing RNE. Frames from Videos 13 and 14. Time in h:min. Scale bar = 10 µm. Movies started at 16 ss-17 ss. (G) Migration speed of rim cells (Mean ± SD). P values for Mann Whitney test with control: laminin Mo p=0.0004, opo Mo p=0.0176, ezrin Mo p<0.0001. See Figure 4—source data 2. |
|
Dynamics of ECM distribution and cell-ECM attachment during RNE morphogenesis. (A) Confocal scans of immunostainings for laminin (middle) and chondroitin sulfate (bottom) in GFP-ras (top) expressing embryos across different stages of RNE morphogenesis. (B) Confocal scans of immunostainings for Fibronectin (top) and phalloidin across different stages of RNE morphogenesis. Merge (bottom). (C) Time-lapse imaging of RNE morphogenesis in embryos expressing Paxillin-mKate2 and GFP-UtrophinCH. Arrowheads point at paxillin enrichment coinciding with UtrophinCH enrichment. Frames are from Video 12. Imaging started at 15 ss. Time h:min. All scale bars = 10 µm. |
|
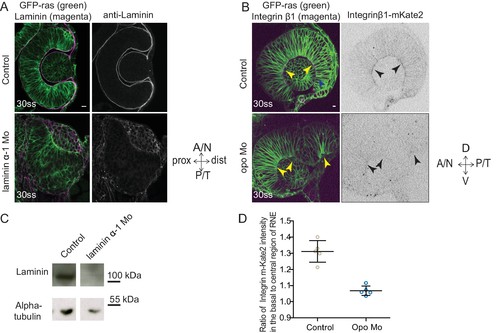
Evaluation of the efficiency of morpholino mediated knockdown of laminin α-1and opo. (A) Confocal scan of 30 ss RNE in control (top) and laminin morphants (bottom) marked with GFP-ras stained for laminin. Morphant shows S-shaped RNE and reduced laminin staining. Scale bar = 10 µm. (B) Confocal scan of side view of 30 ss RNE in GFP-ras and Integrin-β1-mKate2 RNA injected control (top) and opo morphant (bottom) embryo. Morphant shows S-shaped RNE with secondary invagination zone and highly reduced basal localization of integrin. N = 5. Scale bar = 10 µm. (C) Western blot for laminin and alpha-tubulin in control and laminin morphant. (D) Ratio of average integrin β1-mKate2 signal at the basal to central region in the RNE in optical section along the apicobasal axis of the RNE at 30 ss. Mean ± SD, N = 5 embryos. See Figure 4—source data 1. |
|
Effect of perturbed cell-ECM attachment on the optic cup. (A) Confocal scan of 30 ss RNE in control (left) and laminin morphants (right) stained for phalloidin and fibronectin. Morphant shows S-shaped RNE and arrows point at enhanced fibronectin staining. The lower panel shows a zoomed image of the area marked by the red box. N = 6. Scale bar = 10 µm. (B) Confocal scan of 30 ss RNE in control (upper), laminin morphant (middle) and opo morphant (bottom) stained for aPKC. Morphants show S-shaped RNE. Blue arrows point at the apical domain marked by aPKC. Yellow arrowhead points at a few delaminated cells in the laminin morphant. N = 5. Scale bar = 10 µm. (C,D) Normalized average intensity distributions of phalloidin (C) and phosphomyosin (D) in tissue volume along the apicobasal axis of the RNE at 30 ss. Mean ± SEM. Control (brown) n = 25, laminin Mo (green) n = 19 and Opo Mo (blue) n = 20. Tissue sections (n); embryos, N = 5. See Figure 4—source data 3 and 4. |
|
Perturbed basal lamellipodial activity leads to compromised rim involution and impairs RNE morphogenesis. (A) Confocal scan of rim cells in mKate2-ras injected ezrin morphant. Arrow indicates basal blebs in rim cells. Scale bar = 5 µm. (B) Brightfield images of side view of 30 ss optic cup. Control (top) and ezrin morphant (bottom). Arrowhead marks the epithelial accumulation outside the RNE. Scale bar = 50 µm. (C) Confocal scan of optic cup at 30 ss stained for phalloidin. Control (top) and ezrin morphants (bottom). Arrows mark the epithelial accumulation outside the RNE. Asterisk marks the lens. Lookup table indicates the minimum and maximum intensity values. Scale bar = 10 µm. (D,E) Normalized average intensity distributions of phalloidin (D) and phosphomyosin (E) in the tissue volume along the apicobasal axis of the RNE at 30 ss. Mean ± SEM. Control (brown) ezrin Mo (magenta). Tissue sections, n = 25 for control and n = 30 for ezrin Mo; N = 5 embryos each. See Figure 5—source data 1, 2. (F) Time-lapse imaging of RNE morphogenesis in ezrin morphant injected mosaically with ras-mKate2 RNA. Arrows mark rim cells that failed to move. Dashed line marks the outline of developing RNE. Frames from Video 18. Time in h:min. Scale bar = 10 µm. Imaging started at 17 ss – 18 ss. (G) Confocal scan of 30 ss optic cup expressing paxillin-mKate2 and GFP-ras. Control (top), ezrin Mo (bottom). Arrow marks paxillin enrichment at the basal side. N = 7. Scale bar = 10 µm. (H) Confocal scan of paxillin-mKate2 and GFP-ras expressing cells. Control RNE cells (top), ectopic RNE cells in ezrin morphant (bottom). Arrow marks basal paxillin enrichment. Scale bar = 5 µm. |
|
Effect of perturbed cell-ECM attachment on the rim cells. (A) Confocal scan of rim cell in laminin morphant embryos with mosaic expression of mKate2-ras and paxillin-mKate2. Blue arrowhead points at the bleb. Red arrowhead marks at the stable paxillin localization. Frames from Video 15. N = 3. Scale bar = 10 µm. (B) Confocal scan of rim cell in opo morphant embryos with mosaic expression of mKate2-ras and paxillin-mKate2. Blue arrowhead points at the bleb. Red arrowhead marks at the transient paxillin localization. Frames from Video 16. N = 3. Scale bar = 10 µm. (C) Confocal scan of rim cells exhibiting blebs (blue arrowhead) in laminin morphants mosaically labeled by mKate2-ras with DD-myl12b-GFP (left) and GFP-UtrophinCH (right). Yellow arrowhead marks the localization of DD-myl12b-GFP outside and UtrophinCH inside the bleb. Scale bar = 2 µm. (D) Confocal scans of rim zone showing GFP-ras expressing opo morphant clone in control embryos. N = 4 out of 5 transplanted embryos. Blue arrow points at the basal blebs exhibited by the transplanted morphants cells. Scale bar = 5 µm. |
|
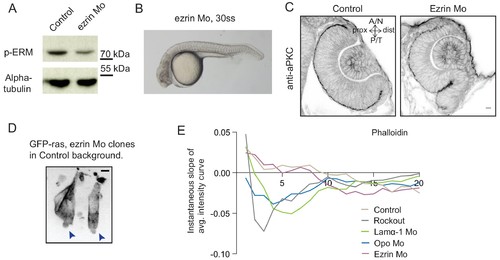
Analysis of ezrin morphant condition. (A) Western blot for p-ERM and alpha tubulin in control and ezrin morphant. (B) Brightfield image of Ezrin morphant embryo at 30 ss stage. (C) Confocal scan of 30 ss RNE in control (left), ezrin morphant (right) stained for aPKC. Morphant shows S-shaped RNE. N = 5. Scale bar = 10 µm. (D) Confocal scan of rim zone showing GFP-ras expressing ezrin morphant clone in a control embryo. N = 2 out of 4 transplanted embryos. Blue arrows point at the basal blebs exhibited by the transplanted morphants cells. Scale bar = 5 µm. (E) Instantaneous slope of the average intensity curves for phalloidin plotted for the basal 20% height of the neuroepithelial volume. The position on the X-axis marks the position along the apicobasal axis. The analysis was performed using the data from Figures 1F, 2C and 5D and Figure 4—figure supplement 3C,D. See Figure 5—source data 3. |
|
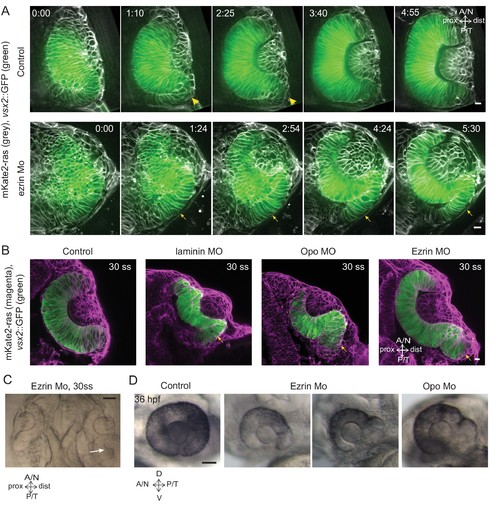
Impairment of rim involution leads to mispositioning of neuroepithelial cells. (A) Time-lapse imaging of control and ezrin morphant condition in Tg(vsx2::GFP, βactin::mKate2ras) background. Arrowheads mark rim zone in control. Arrows mark rim cells that failed to migrate in ezrin morphant. Frames from Videos 19 and 20. Time in h:min. Videos started at 16 ss -17 ss. N = 9. (B) Confocal scans of optic cups at 30 ss in control, laminin morphant, Opo morphant and ezrin morphant conditions in Tg(vsx2::GFP, βactin::mKate2-ras). Embryos were stained for GFP and mKate-2. Arrows point at the ectopic RNE cells. N = 7 each. (C) Brightfield image of 30 ss ezrin morphant showing secondary optic cup phenotype. Arrow marks secondary invagination zone. (D) Brightfield images of 36 hpf control embryos, ezrin morphants and opo morphant. Scale bars in (A,B) =10 µm and (C,D) = 50 µm. |