- Title
-
Development of the cardiac conduction system in zebrafish
- Authors
- Poon, K.L., Liebling, M., Kondrychyn, I., Brand, T., Korzh, V.
- Source
- Full text @ Gene Expr. Patterns
|
The EGFP expression in CCS transgenics (low resolution analysis). A: Confocal Z-stack images of both SqET33-mi28 and SqET33-mi59B in vivo at 30 hpf (I,IV), 48 hpf (II,V) and 72 hpf (III,VI), lateral view. The fluorescence expression in the heart is indicated with a doted circle whereas the cranial ganglia are marked with an arrow and olfactory pit with an arrowhead. EXPRESSION / LABELING:
|
|
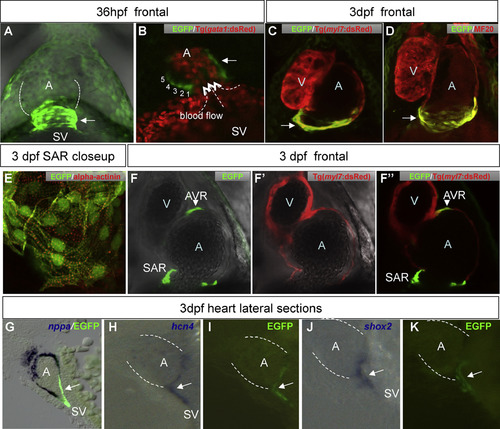
Cardiac EGFP expression in transgenic zebrafish labels the CCS. A: EGFP expression at the venous pole at 36 hpf. sinus venosus(SV) - atrial junction. Numbers indicate rows of EGFP + cells. Dotted arrows indicate direction of blood flow. C-D: Z-stacked confocal images of heart of 3 dpf double transgenic CCS:Tg(myl7:dsRed) (C) and double-labeled EGFP/MF20 (D) shows that the EGFP-positive cells are myocardial in nature. E: EGFP/±-actinin double-labeled EGFP cells at SAR. Image taken at 40× magnifications with 3× digital zoom shows sarcomeric striations (red). F-F′′: Single confocal scan of heart of 3 dpf double transgenic CCS:Tg(myl7:dsRed) shown in EGFP (F), dsRed (F′) and the merged channel (F′′). G: nppa expression does not overlap with the CCS EGFP-signal (sagittal section of 3 dpf heart). H-K: sagittal section of 3 dpf heart of CCS EGFP-positive cells (I,K) co-localize with pacemaker markers hcn4 (H) and shox2 (J). Arrow - SAN EGFP cells, arrowhead - AVC EGFP cells. Dotted white lines in A, H-K outline the atrium. Abbreviations: A, atrium; AVC, atrio-ventricular channel; SAN, sino-atrial node; SV, sinus venosus; V, ventricle. EXPRESSION / LABELING:
|
|
The EGFP expression at the ring of cells at the SAN and AVC is maintained into adulthood in the CCS transgenics. A: A cryosection of a one mpf CCS EGFP + cells stained with anti-acetylated tubulin (red), anti-EGFP (green) and counterstaining with DAPI (blue) shows that the SAR pacemaker cells are neuronally innervated. A′: Enlargement of inset from A - fluorescence at the atrium base. B: In the adult heart, the EGFP expression is localized to the ring of cells located at the SAR and AVR as well as cellular connection between SAN and AVC (labeled by orange arrow). C-C′: Adult whole heart double stained with anti-EGFP (green) and anti-acetylated tubulin (red) shows nerve fibers extending from SAN towards AVC, in close proximity to EGFP cells (green arrows). D: Confocal image of SAN ring showing multiple contacts (blue asterisks) with SAN conduction cells ring. All arrows in this Fig points to SAN EGFP cells whereas arrowheads point to AVC EGFP cells. Dotted white lines in A outline the chambers of the adult zebrafish heart. Abbreviations: A, atrium; AVC, atrio-ventricular region; OFT, outflow tract; SAN, sino-atrial region; SV, sinus venosus; V, ventricle. EXPRESSION / LABELING:
|
|
Loss of endocardium and hemodynamic stimulation affects development of the CCS. A: 3 dpf CCS:clo sibling. B: 3 dpf CCS:clom39-/- mutant. SAN CCS cells (arrows) spread widely in clom39-/-. C, E: Control CCS embryos at 2 dpf and 3 dpf, respectively. D, F: tnnt2 morphants at 2 dpf and 3 dpf, respectively. E′, F′: The same embryos as in E and F, showing dsRed expression of CCS:Tg(myl7:dsRed). Dotted white lines outline the chambers of the embryonic zebrafish heart. A-D: lateral view; E-H: ventral view. Abbreviations: A, atrium; SAN, sino-atrial region; V, ventricle. EXPRESSION / LABELING:
PHENOTYPE:
|
|
One of the three transcripts of fhf2a is cardiac-specific. A: Scheme of the 1 MB genomic region on chromosome 14, where transposons of CCS enhancer trap lines (SqET33-mi28, SqET33-mi59B) are inserted closest to fhf2a/fgf13. zic3, mcf2, atp11c and sox3 are encoded in the reverse strand. B: Gene structure of zebrafish fhf2a highlights its three alternatively spliced variants and transposon insertion sites. The three fhf2a transcripts differ due to differentially spliced first exons, with exon 1S located 2.2 Kb, exon 1U - 6.1 Kb and exon 1Y + 1V - 52 Kb away from shared exon 2. The three transcripts share the common exons 2-5, which encode putative peptides that are 100% identical in amino acid sequence. The ATG (red flag), stop site (black post) and the sizes of introns and exons are indicated. C-F: 1S is the cardiac fhf2a transcript with expression in a SAN at 3 dpf. C: WISH of a 3 dpf embryo with the fhf2a(1S)-specific probe. D,E: Sagittal sections at the level of the 3 dpf heart (D: fhf2a(e2-5)-specific probe; E: fhf2a(1S)-specific probe). White arrows - the base of the heart. F: one step RT-PCR on 3 dpf hearts RNA (top) and brains RNA (bottom). Only the 1S splice variant is expressed in the heart. ef1a - the loading control for PCR. G-N: Distinct expression of fhf2a splice variants as shown by WISH on 3 dpf embryos using isoform specific probes: common exons refer to e2-e5 shared among all three isoforms - G, K; fhf2a(1Y + 1V) - H, L; fhf2a (1U) - I, M and fhf2a (1S) - J, N. Green arrows: dorsal diencephalon; orange arrows: dorsal root ganglia and red arrows. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
|
Reprinted from Gene expression patterns : GEP, 21(2), Poon, K.L., Liebling, M., Kondrychyn, I., Brand, T., Korzh, V., Development of the cardiac conduction system in zebrafish, 89-96, Copyright (2016) with permission from Elsevier. Full text @ Gene Expr. Patterns