- Title
-
Mesp quadruple zebrafish mutant reveals different roles of mesp genes in somite segmentation between mouse and zebrafish
- Authors
- Yabe, T., Hoshijima, K., Yamamoto, T., Takada, S.
- Source
- Full text @ Development
|
Phenotype of the Mesp quadruple homozygous mutant. (A,B) Dorsal view of wild type and Mesp quadruple homozygous mutant at the 1-somite stage. The arrowhead points to the morphological somite boundary. The arrows indicate the notochord. (C-F) Morphology of wild-type (C), mespaa and mespba double mutant (D), mespab and mespbb double mutant (E), and Mesp quadruple homozygous mutant (m4; F) embryos at the 18-somite stage. Magnified images are also indicated (C′-F′). The bracket indicates the impaired somite boundary in the Mesp quadruple mutant. PHENOTYPE:
|
|
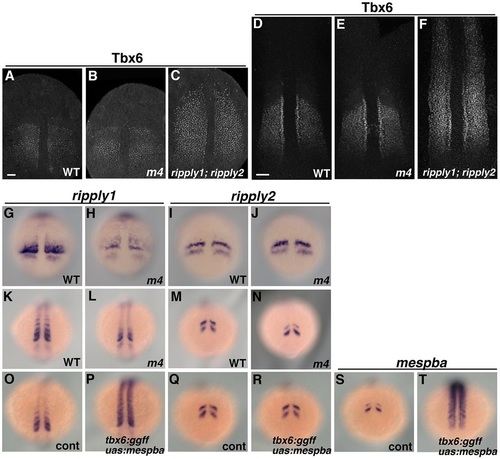
Somite positioning in the Mesp quadruple mutant. (A-F) Immunostaining of Tbx6 in wild-type, Mesp quadruple mutant and ripply1; ripply2 double mutant embryos fixed at the 1-somite stage (A-C) and 9-somite stage (D-F). In wild-type and Mesp quadruple mutant embryos, a clear anterior border of the Tbx6 protein domain is seen. In contrast, in the ripply1; ripply2 double mutant embryo, this domain has expanded anteriorly. (G-N) Expression of ripply1 (G,H,K,L) and ripply2 (I,J,M,N) in wild-type (G,K,I,M) and Mesp quadruple mutant (H,L,J,N) embryos. In the Mesp quadruple mutant, the expression of ripply1 in the anterior PSM is not affected, although expression in the somite region is severely reduced at the 1-somite stage (G,H; 100%; n=15) and 11-somite stage (K,L; 100%; n=14). Expression of ripply2 is not affected at the 1-somite stage (I,J; 100%; n=17) or 11-somite stage (M,N; 100%; n=15). (O-T) The effect of overexpression of mespba on the expression of ripply1 and ripply2 in embryos fixed at the 11-somite stage. In tbx6:ggff; uas:mespba double transgenic fish, the ectopic expression of mespba is detected in the mature somite region and anterior PSM (S,T; n=10; 100%). The expression of ripply1 at the anterior PSM is not affected by the overexpression of mespba, although the expression of somite region is strongly enhanced (O,P; 100%; n=11). The expression of ripply2 is not altered by the overexpression of mespba (Q,R,G,H; 100%; n=7). Scale bars: 50µm. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Impaired formation of the morphological somite boundary in the Mesp quadruple mutant. (A,B) Wild-type (A-A′′) and Mesp quadruple mutant (B-B′′) embryos stained with anti-Fibronectin antibody (A,B) and Phalloidin (A′,B′). The numbers shown in A and B indicate the position of each somite from the most anterior one. White arrows indicate gaps of Fibronectin assembly at somite boundaries. White arrowheads indicate disruption of cell arrangement at a somite boundary. (C,D,G,H) Expression of epha4a in wild-type (C,G) and Mesp quadruple mutant (D,H) embryos. Embryos were fixed at the 1-somite stage (C,D) and the 8-somite stage (G,H). Striped expression of epha4a was lost in Mesp quadruple mutant embryos (D: 100%, n=9; H: 100%, n=12). Black arrowheads indicate striped expression of epha4a in the anterior PSM and somites. (E,F,I,J) Expression of papc in wild-type (E,I) and Mesp quadruple mutant (F,J) embryos. Embryos were fixed at the 1-somite stage (E,F) and the 11-somite stage (I,J). Striped expression of papc was not affected by disruption of the function of the four Mesp genes (F: 100%, n=9; J: 100%, n=18). Scale bar: 50µm. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Impaired rostro-caudal polarity in the Mesp quadruple mutant. Rostral expression of tbx18 (A,B) and fgf8 (C,D) is completely absent in the somites of Mesp quadruple mutant embryos at the 11-somite stage (100%, n=20 and 100%, n=13, respectively). In contrast, caudal expression of uncx4.1 (E,F) and myod (G,H) has expanded in the entire somite of the Mesp quadruple mutant embryo at the same stage (100%, n=20 and 100%, n=18, respectively). The absence of tbx18 expression (I,J) and expansion of uncx4.1 expression (K,L) was also observed at the 28-somite stage (100%, n=15 and 100%, n=22, respectively). Expression of tbx18 is increased (M,N) and that of uncx4.1 is reduced (O,P) in the tbx6:ggff; uas:mespba double transgenic embryo at the 11-somite stage. Magnified images are also shown (M′-P′). The black lines indicate the position of somite boundaries. In the Mesp quadruple mutant at 30dpf (R), hemal arches in several vertebral bones have been duplicated (arrowheads in R′′) and neural arches are split (arrows in R′-R′′′) compared with WT (Q). Magnified images are also indicated (Q′-Q′′′,R′-R′′′). EXPRESSION / LABELING:
PHENOTYPE:
|
|
Abnormal muscle development in the Mesp quadruple mutant. (A-C) Immunostaining of Engrailed (4D9) and Prox1 in the wild type (A) and Mesp quadruple mutant (B). Embryos were fixed at 24hpf. Graph showing the numbers of MPs, identified as the 4D9 and Prox1 double-positive cells, in each somite in the wild-type and Mesp quadruple mutant embryos (C). The number of MPs at 24hpf was counted in the 5-6 somites located in the most posterior part of the trunk region. A total of 16 somites were counted from 3 wild-type embryos and 23 somites from 4 mutant embryos. (D,E) Expression of smyhc in the wild-type and Mesp quadruple mutant embryos at 24hpf. After whole-mount in situ hybridization, the embryos were transversely sectioned at the middle level of the yolk tube. (F,G) Expression of cxcl12a in the wild-type and Mesp quadruple mutant embryos at 24hpf (100%, n=14). (H,I) Expression of pax7 in wild-type and Mesp quadruple mutant embryos in the most posterior region of yolk tube. Embryos were fixed at the 20-somite stage. Scale bar: 50µm. (J-L) Cell-autonomous function of Mesp in superficial horizontal myoseptum formation. (J) Schema for the transplantation experiment. The wild-type cells were transplanted at the marginal zone of Mesp quadruple mutant embryos at the blastula stage. Although the expression of cxcl12a is rescued in the transplanted embryo at 24hpf (K,L), this expression is restricted to the donor cells (K′′,L′′). K and L show the rescued phenotype in different embryos. Although the rescued expression of cxcl12a was observed in a total of 42 cells in 10 embryos, this rescue occurred only in the donor cells in all cases. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Impaired patterning of medial body surface in the Mesp quadruple mutant. In wild-type embryos, the lateral line has been deposited at the medial surface of the trunk and tail region at 2dpf (A). In contrast, the lateral line patterning is impaired in the Mesp quadruple mutant (B). In wild-type embryos, the melanocytes have elongated along the A-P axis and aligned themselves at the medial surface of the trunk and tail at 4dpf (C). In contrast, in the Mesp quadruple mutant embryo, the melanocyte morphology and alignment have been disrupted (D). Unlike the case for the wild-type larva, in which the iridophores are aligned along the mediolateral body surface (E), a gap has formed in the aligned iridophores in the Mesp quadruple mutant at 30dpf (F). PHENOTYPE:
|
|
Normal development of cardiac mesoderm and haematopoietic cells in the mesp quadruple mutant. (A, B) Expression of cmlc2 was not affected by mutations of the 4 mesp genes (100%; n=15). Embryos were fixed at the 17-somite stage. (C, D) Expression of gata1 was not affected by mutations of the 4 mesp genes (100%; n=17). Embryos were fixed at the 11-somite stage. (E, F) Expression of scl was not affected by mutations of the 4 mesp genes (100%; n=15). Embryos were fixed at the 11-somite stage. EXPRESSION / LABELING:
PHENOTYPE:
|
|
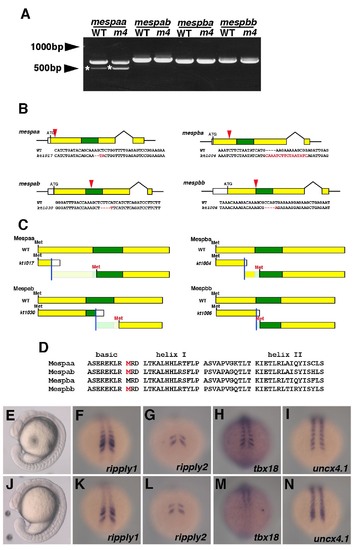
Complete lack of Mesp function in the mesp quadruple mutant (A) RT-PCR products of mesp genes from the wild type and the mesp quadruple mutant embryo. Electrophoresis of mesps cDNA amplified by RT-PCR. The total RNA was isolated from wild type or mesp quadruple mutant embryos at the 8-somite stage. After reaction with reverse transcriptase, cDNA fragment containing the whole sequence of the coding region of th 4 mesps was amplified by PCR. All fragments were confirmed by direct sequencing and any unexpected splicing variant was not detected. Asterisks indicate non-specifically amplified PCR products, checked by direct sequencing. (B-F) Generation of mespabkt1030 and mespbbkt1006 allele. (B) Schematic diagrams showing mutations generated in the 4 mesp genes. Colored boxes indicate the coding regions; especially, green boxes show the basic helix-loop-helix domain. Red arrowheads indicate the approximate position of each mutation. Sequences around the mutation sites are also shown. Red characters in these sequences indicate mutated sequence. (C) Schematic diagrams of predicted protein structures produced from mesp mutant alleles shown in (B). Colored boxes indicate regions where amino acids in the same frame as the original could be translated and green boxes indicate the basic helix-loop-helix domain. White boxes indicate regions where different frames could be translated by frame-shift mutations. Blue lines indicate positions of the mutations. Possible coding frames containing the basic helix-loop-helix domain are also displayed. “Met” colored with red indicate presumptive positions of the translational initiation site. (D) Amino acids sequences of the basic helix-loop-helix domain of the 4 Mesp proteins. Red-colored M correspond to the red-colored Met shown in (C). Note that the presumptive translation products from mutant alleles of mespaa, mespab, and maspbb could not contain the whole basic helix-loop-helix domain. (E,-N) Wild type (E, F, G,H,I) and another mesp quadruple mutant carrying distinct alleles from those mainly examined in this study (J,K,L,M,N). At the 16-somite stage, the morphology of mespaakt1017/kt1017; mespbakt1004/kt1004; mespabkt1030/kt1030; mespbbkt1006/kt1006 was obviously identical to that of mespaakt1017/kt1017; mespbakt1004/kt1004; mespabkt1002/kt1002; mespbbkt1009/kt1009, which were mainly examined in this study (E,F; See Figure 2). The expression ripply1, ripply2, tbx18 and uncx4.1 of mespaakt1017/kt1017; mespbakt1004/kt1004; mespabkt1030/kt1030; mespbbkt1006/kt1006 was also identical those of mespaakt1017/kt1017; mespbakt1004/kt1004; mespabkt1002/kt1002; mespbbkt1009/kt100 at the 11-somite stage (F-I, K-N; See Figure 3 and 5). |
|
Generation of ripply1 and ripply2 double mutant using TALENs. (A) Schematic diagram showing mutations of ripply1 and ripply2. Colored boxes indicate the coding regions of Ripply1 and Ripply2 proteins; green boxes indicate the Ripply-homology domain, which is required for physical interaction with Tbx6, and blue boxes indicate the WRPW motif, which is essential for the interaction with Groucho. Red arrowheads indicate approximate positions of the mutations. The DNA sequences around the mutation sites are given below the schematic diagrams. Red bars indicate the mutated sequence. (B, C) The morphologies of rippley1;ripply2 double mutant embryo at the 13-somites stage. |
|
Generation of tbx6:ggff transgenic fish. (A) Schematic diagram showing construction of the transgene. tol2 and ggff-pA-kan cassette were introduced into CH211-136M16 by using homologous recombination in E.coli. (B,C) The expression pattern of ggff mRNA was almost identical to that of endogenous tbx6 mRNA at the 11-somite stage. (D) The expression of GFP-GFF fusion protein (GGFF) was assessed by immunostaining using anti-GFP antibody. The expression GGFF was detected in the anterior PSM and mature somite region at the 11-somite stage. (E) mcherry expression in tbx6:ggff/uas:gfp-2a-h2a-mcherry double transgenic embryo at the 11-somite stage. Lateral and dorsal views of embryos are shown in B-E and B′-E′, respectively. |