- Title
-
Response to Nodal morphogen gradient is determined by the kinetics of target gene induction
- Authors
- Dubrulle, J., Jordan, B.M., Akhmetova, L., Farrell, J.A., Kim, S.H., Solnica-Krezel, L., Schier, A.F.
- Source
- Full text @ Elife
|
Maternal Smad2 is necessary for mesendoderm specification by Nodal signaling. (A) Illustration of the Smad2 protein showing the position of the ENU-induced non-sense mutation. (B) Western blot against Smad2/3 on 24 hpf embryos of different genotypes for smad2. MZ, maternal-zygotic homozygotes, Z-/-, zygotic homozygotes, Z+/-, zygotic heterozygotes. The pool of maternally contributed Smad2 protein persists for at least 24 hr in zygotic homozygous embryos while it is depleted in MZsmad2 mutants. (C-J) Phenotypic analysis of 36 hpf zebrafish embryos. (C) Wild-type embryo. (D) MZoep embryo: maternal-zygotic mutant for one-eyed pinhead (oep), a cell surface protein required for Nodal signaling (Gritsman et al., 1999). (E) MZsmad2 embryo. Msmad2 mutants display a very similar phenotype (not shown). (F) MZsmad2 embryo rescued with 20 pg of smad2 mRNA. (G-H) MZsmad2 embryo rescued with 50 pg of gfp-smad2 mRNA (brightfield (G), epifluorescence (H)). smad2 mRNA appears to be more effective in rescuing the prechordal plate defects in MZsmad2 mutants as compared to gfp-smad2 mRNA. (I) MZsmad2 embryo injected with 5 pg mRNA for the zebrafish Nodal homolog squint. (J) MZsmad2 embryo injected with 5 pg mRNA for activin. Note that while Activin can activate the Nodal pathway in the absence of oep (Gritsman et al., 1999; Cheng et al., 2003), neither Squint nor Activin can activate the pathway in the absence of Smad2. PHENOTYPE:
|
|
Dynamics of Nodal signaling in vivo. (A) Illustration of Nodal signaling input-output relationship during blastula stage. Gray = yolk, white = blastoderm. Nodal is produced at the margin, diffuses and forms a gradient along the vegetal-animal axis. Nodal signaling induces a gradient of activated Smad2, which induces long- and short-range target gene expression. (B and C) Maximal intensity projection of a confocal stack of a Histone 2B-RFP (B), GFP-Smad2 (C) double transgenic embryo at 50% epiboly (blue box in (A)). GFP-Smad2 strongly accumulates in the nuclei of cells close to the margin, the source of Nodal signals. (D) Heatmap of the nucleo-cytoplasmic (NC) ratio of GFP intensity from the embryo in (B and C). Each dot represents the position of a cell (overlay of five consecutive frames, 3-min intervals per frame). Each cell is color-coded according to its GFP NC ratio (see Figure 2-figure supplement 2 for movement of cells). (E) Examples of single cell tracks at different locations along the vegetal-animal axis, showing changes in GFP-Smad2 NC ratio over time. The position of most cells relative to the margin remains constant during blastula stage. Cells close to the margin activate Nodal signaling earlier and at higher levels than cells at a distance from the margin. The short bursts observed in some cell tracks are caused by transient nuclear accumulation of GFP-Smad2 at the onset of nuclear envelope breakdown and are observed even in the absence of Nodal signaling. (F) NC ratio dynamics of tracked cells along the vegetal-animal axis. (G) Mean NC ratio values from (F) in 30 min bins. Note that the range and amplitude of the Smad2 activity gradient increase over the course of 90 min. Basal NC ratio is higher in younger embryos (see Figure 2G, 3.5 hpf). Since this phenomenon is also observed in the absence of Nodal signaling (MZoep mutants), the higher NC ratio is unlikely to reflect early Smad2 activation, but a higher nuclear import/export ratio of GFP-Smad2 during early development. (H) Time course of ntl (upper panel) and gsc (bottom panel) expression detected by RNA in situ hybridization. ntl begins to be induced as early as 3.5 hpf and its domain of expression expands over time to 100-120 µm from the margin; gsc begins to be induced 30 min later than ntl and its domain of expression expands to 50 µm from the margin. Close-up views of dorsal side, animal pole to the top. Right panel, heatmap for the grayscale intensity of in situ hybridization signals along the vegetal-animal axis showing the increase in range and intensity of ntl and gsc expression over time. See Figure 2-figure supplement 3 for comparison of probes and Figure 2-figure supplement 4 for independent validation of gsc and ntl expression domains using Seurat. |
|
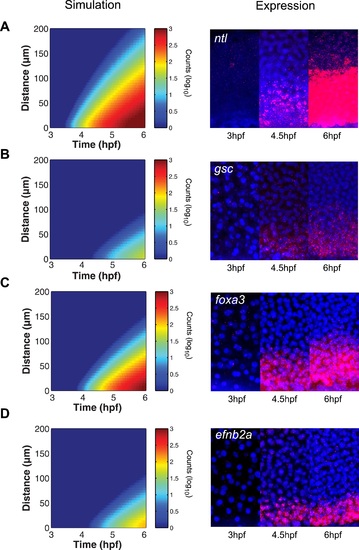
The kinetic model predicts gene expression patterns. Comparison of kinetic model simulations and RNA fluorescent in situ hybridization for ntl (A), gsc (B), foxa3 (C), efnb2a (D). Left panels: simulations of spatiotemporal expression patterns over 3 hr along a 200 µm-high column of cells using gene-specific parameters identified in the parameter screen. Right panels: RNA fluorescent in situ hybridization at 3, 4.5 and 6 hpf. The size of the embryonic field is 100 µm wide and 200 µm high. Animal pole to the top. |