- Title
-
Phagocytosis of mycobacteria by zebrafish macrophages is dependent on the scavenger receptor Marco, a key control factor of pro-inflammatory signalling
- Authors
- Benard, E.L., Roobol, S.J., Spaink, H.P., Meijer, A.H.
- Source
- Full text @ Dev. Comp. Immunol.
|
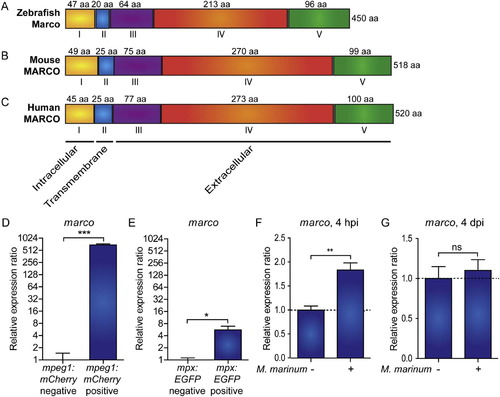
Zebrafish marco encodes a macrophage specific protein with conserved domains and is induced during early M. marinum infection. Comparison of the predicted (A) zebrafish Marco protein structures with (B) murine and (C) human MARCO. The cytoplasmic domain (I), transmembrane domain (II), short spacer domain (III), collagenous domain (IV), and C-terminal cysteine-rich domain (V) are conserved domains detected by SMART analysis. The domains are indicated by Roman numbers; the amino acid (aa) residue lengths of the subunit chains are indicated above their corresponding domains; and the total protein lengths are indicated to the right. The predicted cellular location of each region is indicated below. (D-E) Zebrafish marco is expressed in macrophages. Expression of marco in (D) macrophages and (E) neutrophils. qPCR analysis was performed on RNA from fluorescence-positive cell fractions obtained by cell sorting of 6 dpf larvae transgenic reporter lines for macrophages (Tg(mpeg1:mCherry)) and neutrophils (Tg(mpx:EGFP)). Expression was compared against the fluorescence-negative cell fraction of each transgenic line. (F-G) marco is up-regulated during early M. marinum infection. qPCR analysis was performed on RNA from (F) 4 hpi and (G) 4 dpi mock injected and M. marinum (200 cfu) injected AB/TL embryos. All graphs show data combined from three biological replicates (n = 15 per group, pooled per replicate). |
|
Marco functions in the phagocytosis of M. marinum. Quantification of M. marinum phagocytosis. (A) Schematic overview of the phagocytosis quantification process and representative images of infected control embryos, marco morphants (mo2), and cd36 morphants at 20 mpi. (B–D) Morphants of marco ((B) morpholino 1 and (C,D) morpholino 2) show a delayed phagocytosis of M. marinum (B,C) glycerol stock and (D) 7H9 liquid culture. (E) Morphants of cd36 show no difference in phagocytosis of M. marinum. Each data point represents the percentage of phagocytosed M. marinum in an individual embryo. PHENOTYPE:
|
|
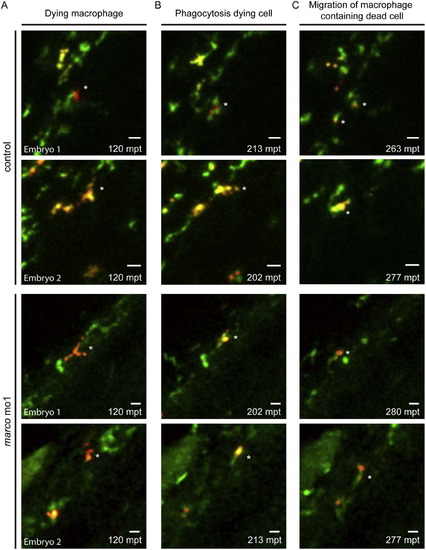
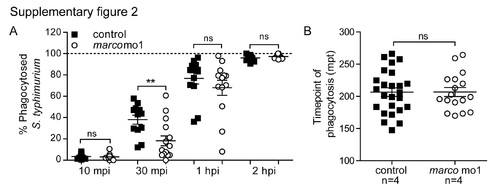
Morphants of marco are capable of phagocytosing dead cells. Phagocytosis of dying cells by macrophages was monitored in time. marco morphants and their controls were imaged after metronidazole treatment which induces nitroreductase-mediated cell ablation in a subpopulation of macrophages (mCherry positive cells). All macrophages express GFP. Time points represented are (A) time of nitroreductase-positive cells undergoing cell death, (B) phagocytosis of the dying cells by healthy GFP-positive macrophages and (C) migration of the macrophage containing the dead cell. Time lapse recordings were made of 4 embryos per group and examples of phagocytosis are shown for 2 control embryos en 2 marco morphants. Quantification of phagocytosis events are shown in Supplementary Fig. S2B. Asterisks indicate the phagocytosing macrophage; mpt: minutes post treatment. PHENOTYPE:
|
|
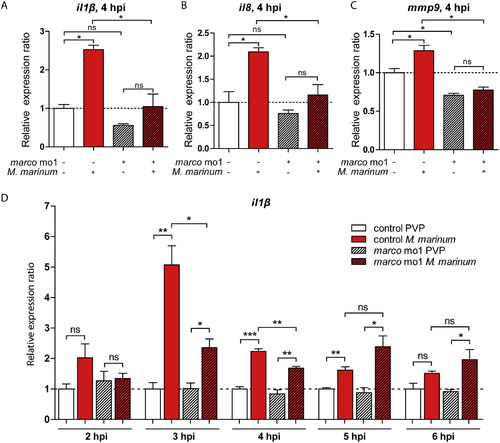
Morphants of marco have suppressed induction of pro-inflammatory cytokine response to M. marinum. (A-C) Effect of marco knockdown on the response of il1b, il8 and mmp9 to M. marinum infection. Expression levels of (A) il1b, (B) il8 and (C) mmp9 were determined by qPCR for marco morphants and their control embryos at 4 hpi after infection with M. marinum (200 cfu) and mock PVP injected controls; (D) Effect of marco knockdown on il1b expression in time (2-6 hpi). Expression was compared against the control PVP sample of each time point. All graphs show data combined from three biological replicates (n = 15 per group, pooled per replicate). |
|
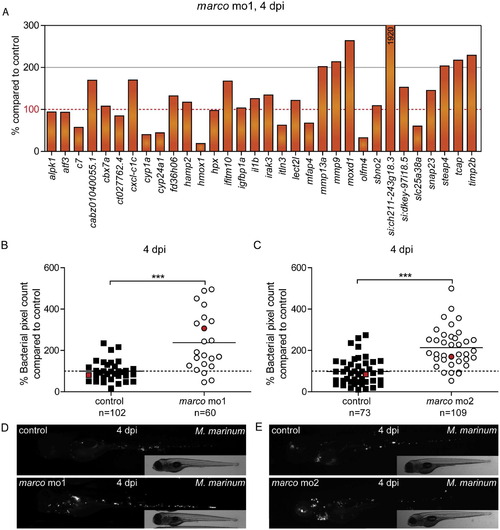
Effect of marco knockdown on the innate immune response and control of M. marinum infection. (A) Graph of the percentage of expression levels of infected marco mo1 morphants compared to infected control embryos on a set of 32 genes that showed reproducible induction by M. marinum infection in control embryos. The expression level of these genes under conditions of marco deficiency is expressed as the percentage of the expression level in the corresponding control. Results are based on RNA-Seq analysis of pools of 30 infected and 30 uninfected embryos for each group. Embryos were injected with M. marinum (200 cfu) or mock injected with 2% PVP, and RNA-Seq analysis was performed at 4 dpi. (B–E) marco knockdown impairs control of M. marinum infection. AB/TL embryos were injected with two different splice blocking morpholinos against (A–D) marco or with control morpholino, subsequently injected with mCherry-expressing M. marinum, and infected embryos were imaged at 4 dpi. (B,C) Bacterial burden was quantified by determining the number of fluorescent bacterial pixels and (D,E) representative stereo fluorescent images are shown below the graph of each experiment. Red symbols indicate which individuals are shown below as representative images. Graphs show combined results of 3 repeated independent experiments. Each data point represents an individual embryo. PHENOTYPE:
|
|
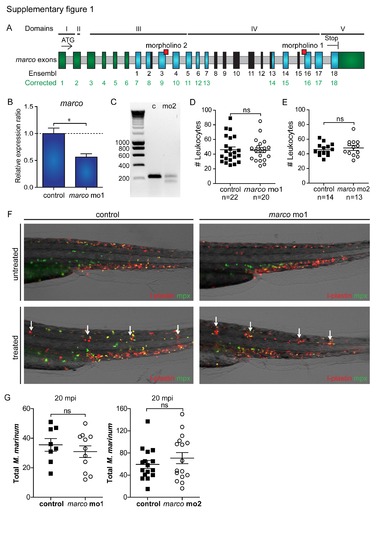
Validation of morpholino knockdown. (A) Corrected annotation of marco exons. The sequenced cDNA clone of marco shows that there are 6 additional exons (shown in green) prior to exon 1 in the current annotation in Ensembl (Zv9, ENSDARG00000059294), which was based on a predicted gene, LOC571584. Furthermore, the cDNA shows partial absence of Ensembl exons 2 and 13 and complete absence of exons 8–12 and 15 (shown in black), and an extended final exon (shown in green). Exons that are not different between the Ensembl annotation and the corrected annotation are shown in blue and introns are shown in grey. Exon–intron boundary sites targeted by morpholinos (red squares), the ATG and stop codon sites, and the conserved protein domains (I to V corresponding with Fig. 1) are indicated. (B,C) Knockdown of marco. The predicted effect of splice blocking morpholinos 1 is deletion of exon 16 in the collagenous domain and the predicted effect of morpholino 2 is deletion of exon 9 in the spacer domain. The exon deletion effect of morpholino 1 could not be confirmed by RT-PCR, but qPCR analysis (28 hpf) demonstrated that the marco mRNA level was effectively reduced (B). Deletion of exon 9 following knockdown of marco with morpholino 2 was confirmed by RT-PCR (C) using RNA from embryos injected with marco morpholino 2 (mo2) or control morpholino (c). The expected RTPCR product size for the control embryos is 228 bp and for marco mo2 morphants is 143 bp. (D,E) Normal leukocyte numbers in marco morphants. marco (D) morpholino 1 and (E) morpholino 2 morphants and control embryos were fixed at 30 hpf and were immuno-labelled with Ab against the general leukocyte marker L-plastin. Leukocyte numbers were counted blinded over the Duct of Cuvier, (ns = not significant). (F) Marco does not affect migration of leukocytes towards local inflammation sites. Representative images of copper sulphate treated control and marco morphant 3 dpf embryos. Embryos were immuno-labelled with Ab against the general leukocyte marker L-plastin (red signal) in combination with a neutrophil-specific Mpx TSA-staining (green signal) at 3 dpf. White arrows indicate accumulation of leukocytes at the local inflammation sites at the neuromasts. (G) Total number of M. marinum counted over Duct of Cuvier at 20 mpi for marco mo1 and mo2 morphants. |
|
Validation of morpholino knockdown. (A) Corrected annotation of marco exons. The sequenced cDNA clone of marco shows that there are 6 additional exons (shown in green) prior to exon 1 in the current annotation in Ensembl (Zv9, ENSDARG00000059294), which was based on a predicted gene, LOC571584. Furthermore, the cDNA shows partial absence of Ensembl exons 2 and 13 and complete absence of exons 8-12 and 15 (shown in black), and an extended final exon (shown in green). Exons that are not different between the Ensembl annotation and the corrected annotation are shown in blue and introns are shown in grey. Exon-intron boundary sites targeted by morpholinos (red squares), the ATG and stop codon sites, and the conserved protein domains (I to V corresponding with Fig. 1) are indicated. (B,C) Knockdown of marco. The predicted effect of splice blocking morpholinos 1 is deletion of exon 16 in the collagenous domain and the predicted effect of morpholino 2 is deletion of exon 9 in the spacer domain. The exon deletion effect of morpholino 1 could not be confirmed by RT-PCR, but qPCR analysis (28 hpf) demonstrated that the marco mRNA level was effectively reduced (B). Deletion of exon 9 following knockdown of marco with morpholino 2 was confirmed by RT-PCR (C) using RNA from embryos injected with marco morpholino 2 (mo2) or control morpholino (c). The expected RTPCR product size for the control embryos is 228 bp and for marco mo2 morphants is 143 bp. (D,E) Normal leukocyte numbers in marco morphants. marco (D) morpholino 1 and (E) morpholino 2 morphants and control embryos were fixed at 30 hpf and were immuno-labelled with Ab against the general leukocyte marker L-plastin. Leukocyte numbers were counted blinded over the Duct of Cuvier, (ns = not significant). (F) Marco does not affect migration of leukocytes towards local inflammation sites. Representative images of copper sulphate treated control and marco morphant 3 dpf embryos. Embryos were immuno-labelled with Ab against the general leukocyte marker L-plastin (red signal) in combination with a neutrophil-specific Mpx TSA-staining (green signal) at 3 dpf. White arrows indicate accumulation of leukocytes at the local inflammation sites at the neuromasts. (G) Total number of M. marinum counted over Duct of Cuvier at 20 mpi for marco mo1 and mo2 morphants. PHENOTYPE:
|