- Title
-
gdf6a is required for cone photoreceptor subtype differentiation and for the actions of tbx2b in determining rod versus cone photoreceptor fate
- Authors
- Duval, M.G., Oel, A.P., Allison, W.T.
- Source
- Full text @ PLoS One
|
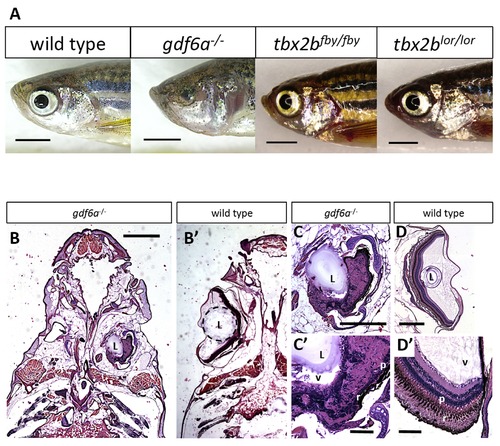
gdf6a and tbx2b mutants do not share the microphthalmic phenotype, despite a shared pathway in early eye development. A. gdf6as327/s327 mutants (labeled gdf6a/ in figures) exhibit microphthalmia to varying degrees of severity during development and throughout adulthood, unlike their wild type and heterozygous siblings. tbx2b mutants do not exhibit microphthalmia, and their eyes develop normally. Scale bars 2 mm. B, C, D. Coronal sections of adult zebrafish heads, comparing microphthalmic gdf6as327/s327 (B) and wildtype fish (B2). Microphthalmia and anophthalmia present variably in gdf6as327/s327 fish (e.g. right and left eyes in B, respectively) and eyes are often noted to possess a lens (L), though in this instance the right eye is inverted such that the anterior segment is oriented towards the midline. RPE (r) and a thin layer of photoreceptors (p) are discernable in gdf6as327/s327 fish (C2), though other retinal layers are not recognizable due to multiple tissue infoldings. In panel D, the lens was presumably displaced away from the iris during dissection/fixation. Note C is at higher magnification compared to D. Scale bar in B 1 mm; C, D is .5 mm; C2, D2 is .1 mm. L, lens; v, vitreous; r, RPE layer; p, photoreceptor layer. PHENOTYPE:
|
|
Disruption of tbx2b does not modify the gdf6a microphthalmic phenotype. A. gdf6as327/s327 mutants exhibit microphthalmia (observed at 3dpf) but tbx2b/ mutants (lor and fby) do not, indicating that disruption of tbx2b does not interfere with identical pathways as gdf6a in early eye development. B. Microphthalmia is rarely observed in tbx2b mutant in-crosses (inx) alone (tbx2b+/lor in-cross shown, n = 220) compared to in-crosses of gdf6a+/s327, which yield 25% with microphthalmia (following Mendelian ratios of inheritance and recessive phenotype). When [gdf6a+/s327;tbx2b+/lor] or [gdf6a+/s327;tbx2b+/fby] compound heterozygous mutants are in-crossed (n = 121 and 195 respectively, both at 6dpf), rates of microphthalmia do not increase significantly from rates expected of in-crosses of gdf6a+/s327 alone (X2 p = 0.873 and p = 0.137, respectively). C. The eye size compared to body length (shown as ratio) of a [gdf6a+/s327;tbx2b+/lor] in-cross does not reveal a subset of intermediate eye sizes, but remains bimodal, with the normophthalmic curve (right curve) showing a normal distribution (Shapiro-Wilk Normality test, W = 0.9888, p = 0.6532) (n = 118, 4dpf). PHENOTYPE:
|
|
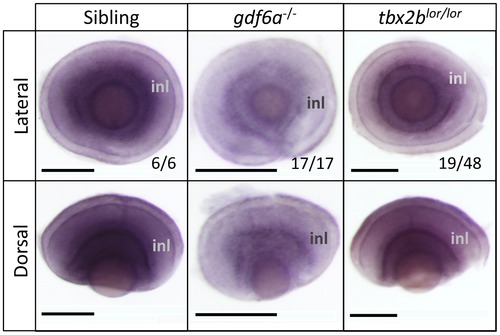
gdf6a positively modulates the abundance of tbx2b transcript during stages of retinal development when photoreceptors differentiate. All panels show in situ hybridization using tbx2b riboprobe. gdf6as327/s327 mutants have less tbx2b expression at 3 days post-fertilization (dpf) compared to normophthalmic siblings, akin to tbx2blor/lor mutants. Fractions represent proportion of clutch represented by image shown. Scale bars 100 μm; inl, inner nuclear layer. EXPRESSION / LABELING:
|
|
Mutation in gdf6a does not disrupt tbx2b function in UV-versus-rod photoreceptor specification, but gdf6a rather plays a role in blue cone specification. A,B. tbx2blor/lor mutants have fewer UV cones and more rods than wildtype fish (the lots-of-rods phenotype) (Kruskall-Wallis ANOVA, **p<0.005), but gdf6as327/s327 mutants have a normal abundance ratio and distribution of UV cones and rods (n = 10 wildtype, 8 tbx2blor/lor, and 7 gdf6as327/s327; UV cones expressing GFP and rods were labeled with antibody 4C12). Scale bars 30 µm and 80 µm, respectively. C. Larval gdf6as327/s327 mutants have a unique cone photoreceptor phenotype in which there are significantly fewer blue cones relative to UV cones at all ages examined (which is not observed in tbx2blor/lor or tbx2bfby/fby mutants- not shown) (Kruskall-Wallis ANOVA, ***p<0.001) Sample sizes at 3 days post-fertilization (dpf) are n = 17 larvae per genotype quantifying cells visualized via opsin in situ hybridization (Panel D); at 4 dpf data are from n = 9 wild type and n = 13 mutants assessed via GFP and mCherry transgene expression in cones; at 6dpf data are from 2 replicates of n = 4+7 wild type and n = 5+6 mutants assessed via transgene expression in cones (Panel E). D. UV and blue cones identified in 3 dpf by in situ hybridization against their respective opsins (Scale bars are 100 µm). E. UV and blue cones identified in transgenic lines at 6dpf by expression of GFP and mCherry, respectively (Scale bars are 60 μm and 40 μm in sibling and mutants, respectively). EXPRESSION / LABELING:
PHENOTYPE:
|
|
A monoclonal antibody raised in rat (10C9.1) labels zebrafish UV cone outer segments, allowing all cone subtypes to be simultaneously labeled by immunohistochemistry. Antibody 10C9.1 specifically labels the outer segments of a class of short single cones in the adult zebrafish retina (Figure S1 panel A). 10C9.1 specificity is supported (see Figure S1 panels B–D), including by a dramatic decrease in number of cells labeled when 10C9.1 is applied to retinas from zebrafish mutants (tbx2blor/lor) that have a paucity of UV cones. A. The population of single cones labeled by 10C9.1 is the UV cones, because established antibodies against the other single cone class, the blue cones, labels a distinct cone population (A2). 10C9.1 enables an unprecedented combination of antibodies raised in different species that simultaneously label and distinguish all cone photoreceptor subtypes (A′′). E. Further evidence that 10C9.1 labels UV cone outer segments comes from its co-localization with UV cones filled with green fluorescent protein (GFP), and its exclusion from blue cones filled with mCherry (mCh) in transgenic zebrafish (Tg(-5.5opn1sw1:EGFP)kj9;Tg(-3.5opn1sw2:m Cherry)ua3011). Panel B is available as Movie S1. Scale bars 30 μm. EXPRESSION / LABELING:
|
|
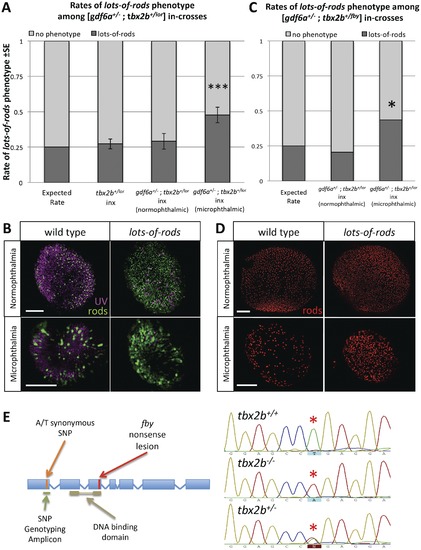
gdf6a modulates tbx2b regulation of UV cone and rod development. A, B. When [gdf6a+/s327;tbx2b+/lor] compound heterozygous mutants are in-crossed (inx), a disproportionate fraction of microphthalmic offspring exhibit the lots-of-rods phenotype compared to normophthalmic siblings, tbx2b+/lor in-crosses, and to predicted Mendelian ratios of the recessive lots-of-rods phenotype (X2 ***p<0.001; 3 replicates of n = 17, 19, 35 microphthalmics; 6dpf). UV cones and rods were labeled using antibodies 10C9.1 and 4C12 displayed in magenta and green, respectively. A portion of microphthalmic larvae with the lots-of-rods phenotype has a tbx2b+/lor genotype (see Table 1). C, D. When [gdf6a+/s327;tbx2b+/fby] compound heterozygous mutants are in- crossed, the lots-of-rods phenotype is again observed at higher rates in microphthlamic eyes compared to normophthalmic eyes (X2 *p = 0.007; 1 replicate, n = 39 microphthalmics, 6 dpf). Panel D shows rod opsin in situ hybridization (red). Scale bars are all 50 μm. E. Genotyping for the lor mutation was performed via linkage analysis using an A/T synonymous SNP located before the DNA binding domain of tbx2b in lor and non-lor alleles, respectively. gdf6as327/s327 mutants with a corresponding SNP of T were used in crossing of the mutant lines. |
|
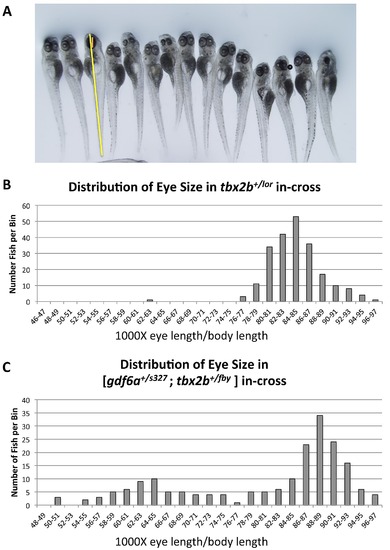
Eye size in various compound mutants shows no obvious change in severity of the microphthalmia phenotype (compare to Figure 1C). A. Eye diameter along the anterior-posterior axis (orange line) was measured at 6dpf and normalized to body length (not including tail fin) (yellow line). Both normophthalmic and microphthalmic larvae are shown. B. Ratios of eye length to body length among the progeny of an in-cross of tbx2b+/lor fish show no obvious difference from wild type fish, (n = 220). C. The same ratios among the progeny of an in-cross of [gdf6a+/s327;tbx2b+/fby] fish show the expected Mendelian abundance of ~25% microphthlamic fish (see also Fig 3B). The normophthalmic fish have eye sizes distributed in a normal fashion (Shapiro-Wilk Normality test, p>0.05). Among the microphthalmic progeny, there is also a normal distribution of eye size (Shapiro-Wilk test, p>0.05) (n = 194). |
|
Antibody 10C9.1 specifically labels the outer segments of a class of short single cones in the adult zebrafish retina, as seen in Figure 5. A. Localization of 10C9.1 labelling to single cone outer segments as clarified by Bodipy counterstain of lipid-rich photoreceptor cell bodies and outer segments. B–D. 10C9.1 specificity is supported by localized labeling in the adult retina (B), a lack of labeling when adjacent retinal cryosections are treated identically except for omission of primary antibody (C), and by a dramatic decrease in number of cells labeled when 10C9.1 is applied to retinas from adult zebrafish mutants (tbx2blor/lor) that have a paucity of UV cones (D). Other negative controls included applying other rat IgGs as primary antibody, and these produced equivalent results to panel C. Retinas in panels B and D were treated identically including equivalent application of 10C9.1 antibody, and simultaneous processing of tissue by inclusion in the same tissue block prior to cryosectioning. The specificity of 10C9.1 is supported by the paucity of labeling in tbx2blor/lor retinas (D), which are known to have few UV cones. Scale bars 30 μm. “rods” indicates rod outer segments; dc, double cones; ipl; inner plexiform layer; onl, outer nuclear layer; inl, inner nuclear layer; rgc, retinal ganglion cell layer. E. An alignment of the antigen used to raise 10C9.1 in rats, which represents the 20 N-terminal amino acids from rainbow trout UV opsin plus a C-terminal cysteine to enable linkage of the peptide to the carrier protein keyhole limpet hemocyanin. |
|
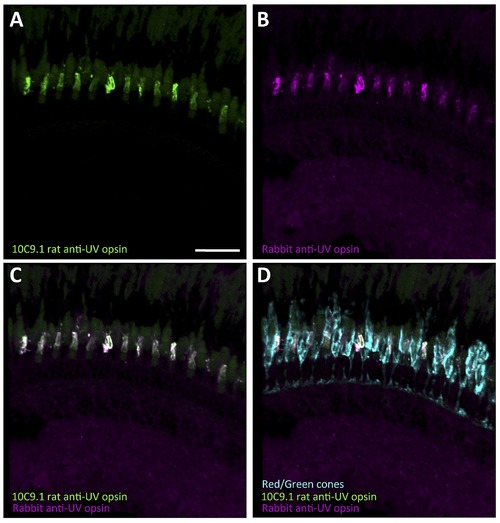
10C9.1 colocalizes with existing rabbit anti-UV antibody (provided by David Hyde, University of Notre Dame). Scale bar 30 μm. Both the 10C9.1 rat anti-UV and Hyde rabbit anti-UV are somewhat over-exposed to demonstrate background/autofluorescent labeling. |