- Title
-
The cone-dominant retina and the inner ear of zebrafish express the ortholog of CLRN1, the causative gene of human Usher syndrome type 3A
- Authors
- Phillips, J.B., Västinsalo, H., Wegner, J., Clément, A., Sankila, E.M., and Westerfield, M.
- Source
- Full text @ Gene Expr. Patterns
|
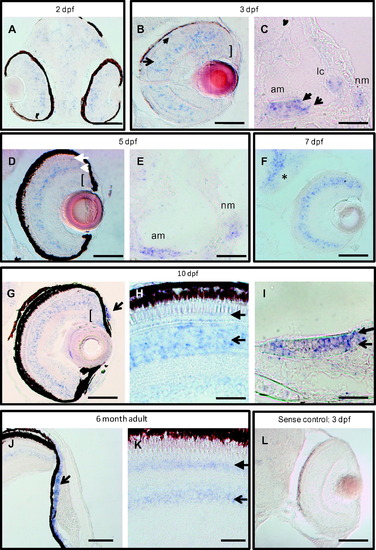
clrn1 is expressed in the retina, inner ear, and lateral line at embryonic, larval, and adult stages. A digoxigenin-labeled cRNA probe was used to detect clrn1 transcript on sectioned tissue from 2 dpf to 6 months of age. (A-C) Developing eyes and ears are enriched for clrn1 in the outer (closed arrow) and inner (open arrow) nuclear layers and ganglion cell layer (bracket) of the retina, (A and B), in mechanosensory hair cells (closed arrowhead in C) and supporting cells (open arrow in C) of the ear and in the neuromast cells (nm). (D-F) Retinal expression is retained through larval stages in all nuclear layers of the retina and in the proliferating cells of the ciliary marginal zone (open and closed arrows in D indicate inner nuclear layer and outer nuclear layer cells, respectively; bracket indicates the CMZ), all sensory patches of ear, and neuromasts (E). Additional expression is noted in brain (asterisk in F). 7 dpf larva in F was raised in PTU to suppress melanocyte formation. (G-I) 10 dpf zebrafish larvae express clrn1 in the anterior chamber of the retina (arrow in G) and continued expression is present in the photoreceptor layer (closed arrowhead in H) inner retina (open arrowhead in H) and CMZ (bracket in G). clrn1 expression is detected both in mechanosensory hair cells (closed arrowhead in I) and supporting cells (open arrow in I) of the sensory patches (anterior macula shown in I). (J and K) Continued expression of clrn1 transcript in the anterior chamber (open arrowhead in J), photoreceptors (closed arrow in K), and inner retinal cells (open arrow in K) of adult retinas. No signal is detected in tissues incubated with a sense probe (shown in L: 3 dpf larval retina treated with PTU). C,E,I: lateral views with anterior to the left; all others horizontal views, with anterior to the bottom. Abbreviations: am: anterior macula; lc: lateral crista; nm: neuromast. Scale bars: A-C, E, H, I and L: 20 µm. D, G, J and K: 50 µm. F: 100 µm. |
|
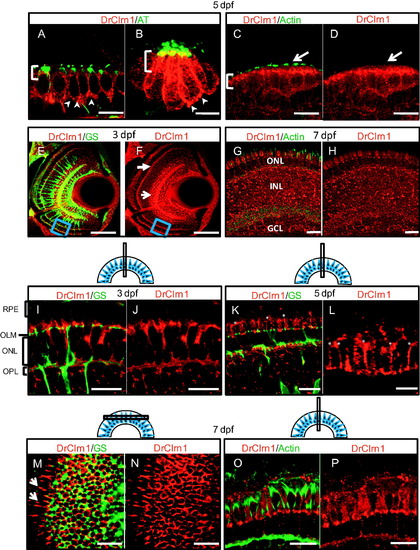
Clarin-1 localizes in mechanosensory hair cell bodies and at the outer limiting membrane and lateral contacts between photoreceptors in the larval retina. (A-D) Cross sections showing Clarin-1 enrichment in apical regions of cell bodies (brackets) and at synapses (arrowheads) of hair cells of the ear (anterior macula shown in A, C) and neuromasts (B). Clarin-1 does not colocalize with actin filaments in the hair cell stereocilia (white arrows in C and D). Kinocilia are labeled with Acetylated Tubulin antibody (AT) and stereocilia are labeled with Actin antibody. (E-H) Cross sections through the retina of 3 dpf (E and F) and 7 dpf (G and H) larvae labeled with DrClrn1 and Glutamine synthetase (GS, panel E) or Actin (panel G) antibodies. Clarin-1 is enriched at synapses (OPL indicated with white closed arrow in F), and in the inner nuclear layer (white open arrow in F) at 3 dpf. Blue box in ventral region of E and F indicates region magnified in panels I and J. A full retina view at 7 dpf shows Clarin-1 label in the photoreceptors (ONL), Inner nuclear layer (INL) and ganglion cell layer (GCL). (I-L) High magnification views of 3 dpf (I and J), and 5 dpf (K and L) retinas. Clarin-1 is enriched at the outer limiting membrane (OLM) and in the outer plexiform layer (OPL). Glial cell processes are seen passing through the layer of the photoreceptor nuclei (ONL). At 3 dpf, the apical structures of photoreceptors, the inner and outer segments, are rudimentary and the retinal pigmented epithelium (RPE) lies close to the OLM. By 5 dpf, cone apices have extended such that space between the OLM and the RPE has increased, and fine, filamentous enrichments of Clarin-1 can be observed between cone inner segments (asterisks in K and L; high magnification image shown in L). (M and N) Transverse section through the 7 dpf retina to visualize the photoreceptor layer from the top down. The meshwork of cell junctions that make up the OLM derive from Müller cell processes labeled by Glutamine synthetase (GS). Clarin-1 localization partially overlaps, and is also enriched at the lateral interfaces extending from the periphery of the transverse cut (white arrows in I). (O and P) Clarin-1 is seen in close proximity to actin at the lateral contacts between photoreceptors in the 7 dpf retina. Scale bars: E, F: 50 µm; A, C, D, G-P: 10 µm. B and L: 5 µm. Schematics show the plane of sectioning for retinal tissue. EXPRESSION / LABELING:
|
|
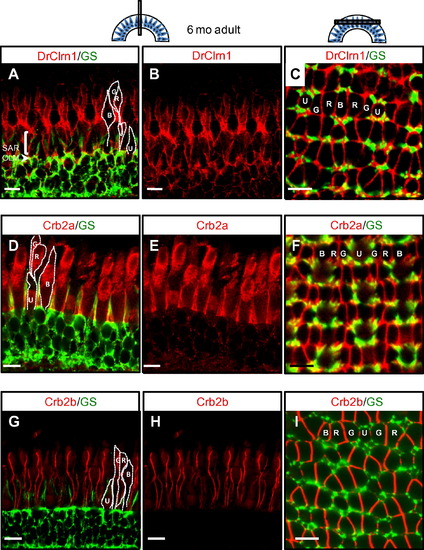
Clarin-1 localizes at interfaces between photoreceptor inner segments in the adult retina. (A-C) Cross (A and B) and transverse (C) sections through adult retinas show Clarin-1 localization partially overlapping with glial cells labeled with Glutamine Synthetase (GS) at the OLM (arrowhead). Enrichment of Clarin-1 is also noted in the subapical region (SAR) and between all cone inner segments in the outer retina. The silhouettes of the four cone subtypes are outlined in white in A, D and G and the mosaic cone arrangement is labeled in C, F and I. (D-I) Cross (D-E, G-H) and transverse (F, I) sections of adult retinas labeled with Crb2a (D-F) and Crb2b (G-I) antibodies recapitulate results published by Zou et al., 2012, and provide points of reference for the Clarin-1 (A-C) localization pattern at lateral cone interfaces. Abbreviations: OLM: outer limiting membrane; G: green opsin cone, R: red opsin cone; B: blue opsin cone; SAR (subapical region) U: UV opsin cone. Scale bars: 10 µm. EXPRESSION / LABELING:
|
Reprinted from Gene expression patterns : GEP, 13(8), Phillips, J.B., Västinsalo, H., Wegner, J., Clément, A., Sankila, E.M., and Westerfield, M., The cone-dominant retina and the inner ear of zebrafish express the ortholog of CLRN1, the causative gene of human Usher syndrome type 3A, 473-81, Copyright (2013) with permission from Elsevier. Full text @ Gene Expr. Patterns