- Title
-
Transcriptional signature of accessory cells in the lateral line, using the Tnk1bp1:EGFP transgenic zebrafish line
- Authors
- Behra, M., Gallardo, V.E., Bradsher, J., Torrado, A., Elkahloun, A., Idol, J., Sheehy, J., Zonies, S., Xu, L., Shaw, K.M., Satou, C., Higashijima, S.I., Weinstein, B.M., and Burgess, S.M.
- Source
- Full text @ BMC Dev. Biol.
|
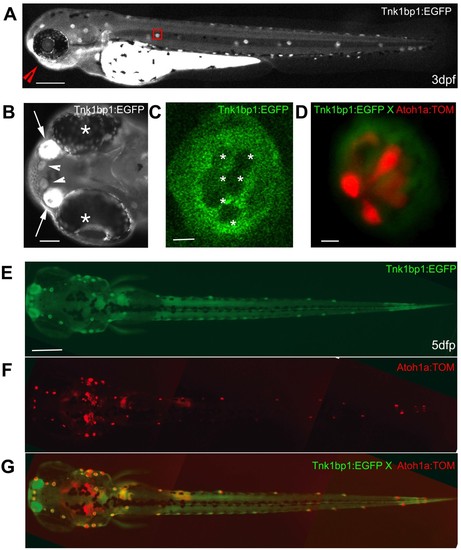
Live imaging of a 3 day old Tg (tnks1bp1:EGFP) embryo and a 5 day old Tg(tnks1bp1:EGFP) x Tg (atoh1a:dTOM) double transgenic larva. A. GFP is expressed in all of the neuromasts in the anterior (head) and posterior (trunk and tail) lateral line, as shown in a three day old embryo. B. Ventral view (red arrowhead in A) of the rostral head region of the embryo, showing a strong GFP expression in the olfactory epithelia (white arrows) between the eyes (white asterisks) and in the two more rostral neuromasts (white arrowheads). C. Magnification of a neuromast (red box in A) in a 3 dpf live embryo, showing GFP expression excluded from the six centrally located hair cells (white asterisks). D. Neuromast of a Tg(tnks1bp1:EGFP) × Tg(atoh1a:dTOM) 5 dpf double transgenic animal expressing GFP (green) in the accessory cells and TOMATO (red) in hair cells. E, F and G. Dorsal view of a 5 day dpf Tg(tnks1bp:GFP) × Tg(atoh1a:dTOM) double transgenic larva. - 200 microns in A and E, 40 microns in B, and 5 microns in C and D. EXPRESSION / LABELING:
|
|
Immuno-staining in neuromasts of 5 dpf Tg (tnks1bp1:EGFP) larvae. A. B. Confocal images of two different neuromasts (1 and 2) double-stained with anti-GFP (green, first and last columns) and anti-Myosin VI (red, second and last columns). The GFP positive cells (green) are the accessory cells (i.e. a combination of supporting cells and mantle cells). Around the apical pole of hair cells, accessory cells form a precisely organized honeycomb like annular structure (white arrows in A and B, first columns). Hair cells send their tightly packed hair bundles through the openings in the top, (white arrowheads in A, middle panel). C. Schematic cross-section (left image) and dorsal view (right image) of a neuromast illustrating the respective position of the accessory cells (cytoplasm light green and nucleus dark green) and the hair cells (cytoplasm dark red and nucleus light red). D. Cryosection of a neuromast, immunolabeled with an anti-GFP antibody. GFP expression is excluded from all hair cells (nucleus and cytoplasm) and found in all accessory cells, comprising the supporting and mantle cells (white stars). - 10 microns in A and B, 5 microns in D. EXPRESSION / LABELING:
|
|
The trapped gene is a homolog of the tnk1bp1 gene. A. The tnks1bp gene. It encodes 8 exons with the first one (black square), being untranslated. The gene-trap construct landed in the first intron (green arrow), resulting in a product described in (C). Two sets of primers for qRT-PCR were designed to track the presence of alternatively spliced products in transgenic larvae. The amplified products are marked by the two colored lines under the gene depiction. The lines illustrate respectively: red for the product comprised of exon1 (black square) to the gfp from the gene-trap (not depicted), blue for exon1 (black square) to exon2 (blue square of the tnks1bp1 gene. B. Alignment of a stretch of ~70AA of a portion of the C-term of the putative TNKS1BP1 product with ten partial sequences of the putative product of the TNK1BP1 as sequenced/predicted from 10 species. The amino acids highlighted in green are identical and the ones in blue are conservatively substituted. C. Symbolic representation of the mRNA, which was found in abundance in the transgenic embryos. This messenger results from the fusion of the first untranslated exon 1 of the tnks1bp1 gene and the gpf sequence followed by a polyA tail, elements encoded by the integrated transposon construct (green arrow in A). D. Three tail neuromasts are positive (white arrows) by in situ hybridization with a probe against the tnks1bp1 gene. D. and E. Neuromasts after WISH (against tnks1bp1), showing stronger hybridization in the more external cells, corresponding to accessory cells. EXPRESSION / LABELING:
|
|
Strategy to establish the transcriptional profile of accessory cells combining FACS and microarrays. A. Schematic of the experimental procedures. B. illustration of the gating used for the FACS. The left plot (showing the P1 gate) was used to sort cells according to cell size (forward scatter, FSC-A) vs. granularity (side scatter, SSC-A). The right plot discriminated cells according to GFP fluorescence intensity (GFP FITC-A) vs. Phycoerythrin (PE-A). Gates were selected as shown to sort GFP negative (GFP Neg) and GPP positive (GFP Pos) cells. |
|
WISH and q-RT-PCR on novel genes enriched in accessory cells A to K. Whole mount in situ hybridizations (WISH) in 3 and 5 dpf larvae A ring like staining is visible, because hybridization is excluded from centrally located hair cells and only found in peripheral accessory cells for gene #9, 10, 11 and 12 (A to G). In #6 and 8 (H to K), the hybridization is present in the whole neuromast, but appears much stronger in centrally located hair cells. A. WISH against #9 in 3 three neuromasts in a 3 dpf larva (A). B and C. WISH against #10 in a neuromast in a 3 dpf (B) and a 5 dpf larva (C). D and E. WISH against #11 in a portion of the 5 dpf larval head showing 4 neuromasts (D) and of a neuromast in a 3 dpf larva (E). F and G. WISH against #12, in two neuromasts in 5 dpf larvae (F and G). H and I. WISH against #6 in a neuromast in a 3 dpf (H) and in a 5 dpf larvae (I). J and K. WISH against #8 in two neuromasts in 5 dpf larva (J and K). - is 50 microns in A and D, and 5 microns in B. L. Average fold increased expression of genes determined by qRT-PCR. The tnks1bp1 gene is in green. Genes showing an expression pattern by WISH (in A to Q) are in yellow the others are in blue. Error bars represent standard deviation. EXPRESSION / LABELING:
|

Unillustrated author statements EXPRESSION / LABELING:
|