- Title
-
Alcama mediates Edn1 signaling during zebrafish cartilage morphogenesis
- Authors
- Choudhry, P., Joshi, D., Funke, B., and Trede, N.
- Source
- Full text @ Dev. Biol.
|
alcama morphants have defects in facial skeletal patterning. (A) Graphic representation of alcama RNA illustrating the position of TB and SB MOs. (B) Nomarski image of a 48 hpf WT larva with the box indicating the region shown in (C)–(E). 48 hpf control (C), alcama TB (D), and alcama SB (E) morphants stained with Zn-5 antibody. Both MOs effectively knock down alcama expression in the heart (arrowheads) and pouches (arrows). (F, G) Lateral views of 5 dpf larvae injected with control and alcama TB MOs. alcama morphants have a protruding jaw (arrows), cardiac edema (arrowheads) and an absent swim bladder (open arrows). Lateral views of Alcian blue-stained control (H) and alcama (I) morphants at 5 dpf. The line indicates the length of the pharyngeal cartilage, which is shortened in alcama morphants. The arrow points to Meckels cartilage, which is bent ventrally in alcama morphants. Scale bars: 50 μm. |
|
alcama morphants have cartilage defects similar to the edn1 class of mutants. Flatmounts of mandibular and hyoid cartilage from 5 dpf Alcian Blue-stained larvae (A, C, E, G); corresponding schematics (B, D, F, H). The joint between Meckel′s and palatoquadrate is fused in alcama morphants and furinA-/- and edn1-/- mutants. Similarly, the interhyal is absent in alcama morphants and furinA-/- mutants leading to a fusion of the ceratohyal and hyosymplectic cartilages. alcama morphants and furinA-/- mutants have misshapen Meckel′s cartilage and ceratohyal, but edn1-/- mutants have the most severe defect with a lack of ceratohyal and severely misshapen Meckel′s cartilage. DV joint regions are indicated with arrows in A and B. Fusions at joints are indicated with asterisks in (C)–(H). Cartilages are labeled as followed: pq (palatoquadrate), mc (Meckel′s cartilage), hm (hyomandibula), ch (ceratohyal), sy (symplectic), and ih (interhyal). Scale bar: 50 μm. PHENOTYPE:
|
|
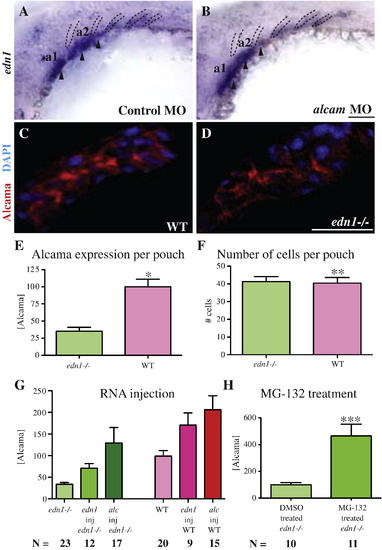
Edn1 regulates Alcama levels. edn1 is expressed in the mesodermal cores (arrowheads) of the first three arches and in pharyngeal pouches 2–4 (dotted lines) (A). Its expression is unchanged in alcama morphants (B). Cropped image of a single pharyngeal pouch from 30 hpf WT sibling (C) and edn1-/- mutant (D) stained with Zn-5 (anti-Alcama) antibody in red and DAPI in blue. (E) Bar graph showing the difference in total Alcama protein in a pouch (second or third pouch), represented as measured fluorescence intensity normalized to WT; *p-value < 0.0001 by a two-tailed t-test. The bar graph in (F) shows the total number of cells per pouch is unchanged in WT and edn1-/- mutants; **p-value = 0.08495 by a two-tailed t-test. n = 14 for WT; n = 11 for edn1-/- mutants for this experiment which was repeated with similar results. (G) edn1 regulates the Alcama levels in pouches. The bar graph shows the sum of Alcama intensity in the first three pouches of 30 hpf edn1-/- (green) and WT (red) larvae after the indicated injections (p-value < 0.0001 by one-way analysis of variance). (H) Bar graph showing the sum of Alcama intensity in the first three pouches of 30 hpf edn1-/- larvae after treatment with DMSO or the proteasome inhibitor MG-132; ***p-value = 0.0009 by a two-tailed t-test. N is the number of larvae in a single experiment, which was repeated with similar results. a1 and a2 label pharyngeal arches 1 and 2. Scale bars: 50 μm. EXPRESSION / LABELING:
|
|
alcama is required for edn1-dependent gene expression. Ventral views of larvae at 30 hpf (A–C), and lateral views at 30 hpf (D–L) and at 48 hpf (M–O) respectively. ISH for hand2 (A–C), dlx3b (D–F), dlx5a (G–I), dlx6a (J–L), and bapx1 (M–O) in control and alcama morphants and in edn1-/- mutants. hand2, dlx3b, dlx5a and dlx6a expression is strongly reduced in the first and branchial arches and moderately reduced in the second arch in alcama morphants. Their expression is strongly reduced in all arches in the edn1-/- mutants. bapx1 expression in the first arch intermediate domain is moderately reduced in alcama morphants and strongly reduced in edn1-/- mutants. a1 and a2 label pharyngeal arches 1 and 2. Dotted lines highlight pharyngeal pouches 1 and 2. Scale bars: 50 μm. EXPRESSION / LABELING:
|
|
alcama rescues NC differentiation defect in edn1-/- mutants. Lateral views of 30 hpf larvae with representative absent (A), partial (B) or strong (C) staining for dlx5a. Bar graphs show the percentage dlx5a expressing larvae in edn1-/- mutants (D) and WT (E) following the stated injections. edn1 or alcama RNA was injected with or without alcama MO. alcama RNA decreases the percentage of affected individuals (absent dlx5a staining) by nearly 30% in edn1-/- mutants and co-injection of alcam Mo abrogates the ability of edn1 RNA to rescue dlx5a expression in edn1-/- mutants. Indicated p-values are calculated by Fishers exact test. n depicts the number of experiments and N depicts the total number of larvae represented in the columns of the plot (p-value < 0.0001 for both D and E by χ2 analysis). a1 and a2 label pharyngeal arches 1 and 2. Dotted lines highlight pharyngeal pouches 1 and 2. Scale bar: 50 μm. |
|
nadl1.1 morphants phenocopy alcama morphant cartilage defects. Dorsal views (A–C) and lateral views (D, E) of 30 hpf larvae. nadl1.1 is expressed in the pharyngeal arches, diencephalon, telencephalon, hindbrain neurons, neural tube and pectoral fin in 30 hpf WT larvae (A). 30 hpf nadl 1.1 morphants have down-regulated hand2 (C) and dlx5a (E) expression as compared to uninjected WT (B, D). (F) Bar graph showing that the percentage of larvae with down-regulated (absent) dlx5a expression remains unchanged in the p53-/- background (p-value = 1.000 by Fishers exact test, indicating that WT and p53-/- mutants are not significantly different). Dissected mandibular and hyoid elements from 5 dpf WT and nadl1.1 morphant larvae stained with Alcian blue (G, H) and their corresponding schematics (G′, H′). The joint fusions in nadl1.1 morphants (H, H′) marked by * resemble those seen in alcama morphants. (I) Bar graph comparing the percentage of larvae with affected cartilage in alcama morphants in WT versus p53-/- mutants (p-value = 0.1941 by Fishers exact test, indicating that WT and p53-/- mutants are not significantly different). N is the total number of larvae from three experiments. Scale bars: 50 μm. |
|
alcama and nadl1.1 interact during cartilage formation. (A) Lateral view of a 30 hpf WT larva stained with Zn-5 antibody (Fast Red) and anti-sense nadl1.1 RNA (Fluorescein), revealing that nadl1.1 is expressed in the NC adjacent to alcama in the endoderm. The bar graphs display the percentage of 30 hpf edn1-/- (B) and WT (C) larvae with dlx5a staining after the indicated injections. (B) shows that nadl1.1 is required for alcama rescue of dlx5a expression in edn1-/- mutants (p-value < 0.0001 by Fishers exact test). (C) and (D) show that alcama and nadl1.1 interact during dlx5a expression and cartilage morphogenesis. (D) Bar graph displaying the percentage of 5 dpf larvae with affected cartilage after the indicated injections (p-value < 0.0001 by χ2 analysis). n indicates the number of experiments and N the total number of larvae. Scale bar: 50 μm. EXPRESSION / LABELING:
|
|
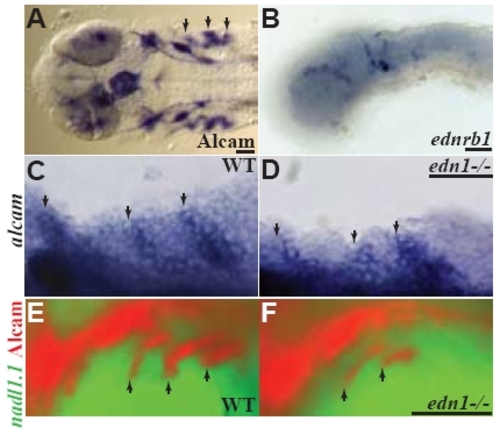
alcama and nadl1.1 ISH in edn1-/- mutants. (A, B) 30 hpf WT larvae showing expression of alcama and ednrb1 respectively. (C, D) alcama ISH showing expression in pharyngeal pouches of WT and edn1-/- larvae at 30 hpf. (E, F) 30 hpf larvae stained with Alcama antibody (Fast Red) and nadl1.1 RNA (Fluorescein). nadl1.1 expression in the NC is unchanged while Alcama is down-regulated in edn1-/- mutants (F). Arrows point to alcama expression in the pharyngeal pouches. Scale bars: 50 μm. EXPRESSION / LABELING:
|
|
edn1-mediated rescue of Alcama expression. Representative images of the data shown in Fig. 3G. Cropped images of pharyngeal pouches 1-3 from 30 hpf larvae stained with Zn-5 antibody and photographed at the same exposure. Alcama expression in edn1-/- mutants (A) is rescued to near WT levels (C) upon injection of edn1 RNA (B). Scale bar: 50 μm. |
|
NC and endoderm are patterned and specified correctly in alcama morphants. 30 hpf control (A,C,E) and alcama morphants (B,D,F). Tg(fli1:EGFP) larvae stained with Zn5 antibody in red (A,B) show that NC is patterned correctly in absence of alcama. dlx2a (C,D) and nkx2.3 (E,F) expression in NC and endoderm, respectively, reveals correct specification of NC and endoderm. a1 and a2 label pharyngeal arches 1 and 2. Dotted lines highlight pharyngeal pouches. Scale bars: 50 μm. |
|
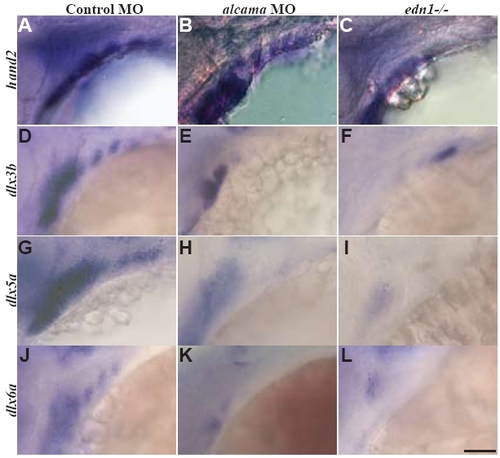
hand2 expression recovers by 48 hpf in alcama morphants. Lateral views of 48 hpf control and alcama morphants and edn1-/- mutants showing hand2 (A-C), dlx3b (D-F), dlx5a (G-I), dlx6a (J-L) expression. dlx3b, dlx5a and dlx6a expression is strongly reduced in the first and branchial arches and moderately reduced in the second arch in alcama morphants. However hand2 expression recovers to WT levels in alcama morphants (B). Scale bar: 50 μm. |
|
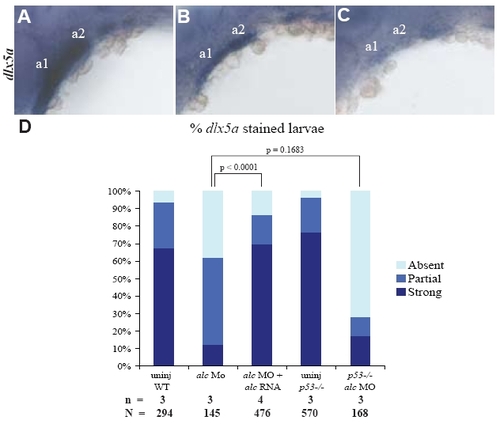
NC differentiation defect is caused by alcama knockdown. 30 hpf larvae showing strong (A), partial (B) and absent (C) dlx5a expression. (D) Bar graph showing the percentage of 30 hpf larvae in each of the three categories of dlx5a expression. alcama RNA decreases the percentage of affected individuals (sum of absent and partial dlx5a staining) in alcama morphants to WT levels (p-value < 0.0001 by Fishers exact test). The percentage of affected individuals is similar in WT and p53-/- (p-value = 0.1683 by Fishers exact test). n indicates the number of experiments and N the total number of larvae. |
Reprinted from Developmental Biology, 349(2), Choudhry, P., Joshi, D., Funke, B., and Trede, N., Alcama mediates Edn1 signaling during zebrafish cartilage morphogenesis, 483-493, Copyright (2011) with permission from Elsevier. Full text @ Dev. Biol.