- Title
-
Tissue-specific requirements for specific domains in the FERM protein Moe/Epb4.1l5 during early zebrafish development
- Authors
- Christensen, A.K., and Jensen, A.M
- Source
- Full text @ BMC Dev. Biol.
|
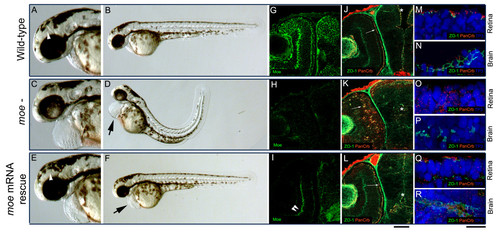
Injection of moe mRNA rescues defects in moe- embryos. (A-B) At 60 hpf, in wild-type embryos, the floor of the diencephalic ventricle is visible (A, white arrow head), the RPE is uniform, and the body axis is straight. (C, D) In moe- embryos, the ventricles are small or absent, the RPE is patchy, the tail curves and there is pericardial edema (D, arrow). (E, F) In moe- embryos injected with moe mRNA, the floor of the diencephalic ventricle is visible (E, white arrow head), the RPE is uniform, and the body axis is straight but mild pericardial edema persists (F, arrow). Anti-Moe labeling of 60 hpf wild-type embryos (G), moe- embryos (H), and moe- embryos injected with moe mRNA (I): the plexiform labeling in moe- embryos injected with moe mRNA (I, double arrowheads) is largely background. Adherens junctions (ZO-1, gnjected with moe mRNA (I): the plexiform labeling in moe- embryos injected with |
|
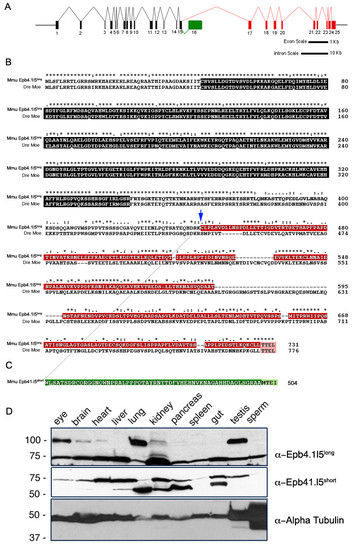
Genomic structure of the mouse epb4.1l5 locus and expression of its two major splice isoforms. (A) Diagram of the inton/exon structure of the Mus musculus epb4.1l5 locus. Exons that are common to both isoforms are black, the exons unique to epb4.1l5long are indicated in red and the unique exon in epb4.1l5short is indicated in green. Bars represent 1 kb and 10 kb scales for exon and intron lengths, respectively. (B) ClustalX alignment of mouse Epb4.1l5long and zebrafish Moe. Ymo1long and Moe share a high degree of homology within the FERM domain (black). Ymo1long and Ymo1short are identical up to Lysine 444 (blue arrow) and then alternately spliced into the long (red) and short (green) isoforms. Moe and Epb4.1l5long, and Epb4.1l5short have predicted C′terminal PDZ-binding domains (Pink [TTEL]) and (light green [MTEI]). (*) identical, (:) highly conserved, (.) Moe and Epb4.1l5long, and Epb4.1l5short have predicted C′terminal PDZ-binding domains (Pink moderately conserved. (C) Western analysis of mouse tissues with antibodies raised against the unique C′terminal sequences of Epb4.1l5long and Epb4.1l5short. Two bands are immunoreactive with the anti-Epb4.1l5long antibody, one migrates at the expected molecular weight of Epb4.1l5long (100 kDa) and is present in the eye, brain, heart, lung, kidney, and testis. An additional band migrates at 75 kDa, which is probably non-specific. Two bands are recognized by the affinity purified Epb4.1l5short antibody. A lower band migrates at the predicted molecular weight of 56 kDa and is present in brain, liver, lung, kidney, pancreas, and gut. A second band with a broad expression pattern migrates at approximately 75 kDa. Anti-α-Tubulin was used as a loading control. |
|
Both the PDZ-binding domain and unique sequences in Epb4.1l5long required for rescue of moe- mutant defects. (A-C) At 30 hpf in wild-type embryos, Moe localizes cortically in brain and retinal neuroepithelial cells and is concentrated at the apical surface (A) and ZO-1 (green) and panCrb (red) localize to the apical surface of the retina (B) and brain (C). (D-F) At 30 hpf in moe- embryos, there is no Moe labeling (D) and ZO-1 (green) and panCrb (red) fail to localize to the apical surface of the retina (E) and brain (F). (G) At 30 hpf in moe- embryos injected with epb4.1l5long mRNA, Epb4.1l5long immunoreactivity is cortically localized in most retinal and brain neuroepithelial cells and ZO-1 (green) and panCrb (red) localize to the apical surface in the retina (H) and brain (I). Upper inset (G), magnified section of anti-Epb4.1l5long labeling and lower inset, uninjected moed ZO-1 (green) and panCrb (red) localize to the apical surface in the retina (H) and brain (I). Upper inset (G), magnified sect- embryos shows no labeling with anti-Epb4.1l5long. (J) At 30 hpf in moe- mutants injected with epb4.1l5short mRNA, anti-Epb4.1l5short is cytoplasmically localized and ZO-1 (green) and panCrb (red) do not localize to the apical surface in moe- retina (K) and brain (L). Upper inset (G), magnified section of anti-Epb4.1l5short labeling and lower inset, uninjected moe- embryos shows no labeling with anti-Epb4.1l5short. (M, N) At 30 hpf in moe- mutants injected with myc-epb4.1l5FERM mRNA, ZO-1 (green) and panCrb (red) do not localize to the apical surface in moe- retina (M) and brain (N). (O, P) At 30 hpf in moe- mutants injected with myc-epb4.1l5long_ΔPBD mRNA, ZO-1 (green) and panCrb (red) do not localize to the apical surface in moe- retina (O) and brain (P). (Q, R) At 30 hpf in moe- mutants injected with myc-epb4.1l5short+long_PBD mRNA, ZO-1 (green) and panCrb (red) localize to the apical surface in moe- retina (Q) and brain (R). (S, T) At 30 hpf in moe- mutants injected with myc-epb4.1l5long+short_PBD mRNA, ZO-1 (green) and panCrb (red) localize to the apical surface in moe- retina (S) and brain (T). (U) Western analysis of zebrafish embryos injected with epb4.1l5long and epb4.1l5short mRNA were tested for expression of protein product with isoform-specific antibodies at time points from 6 hpf to 96 hpf. (V) Western analysis of zebrafish embryos injected with epb4.1l5FERM (anti-Myc), epb4.1l5long_ΔPBD (anti-Epb4.1l5long), epb4.1l5short+long_PBD (anti-Myc), epb4.1l5long+short_PBD (anti-Epb4.1l5long). Blots stripped and reprobed with Anti-α-Tubulin as a loading control. Scale bars, 10 μm (A, D, G, J), 50 μm (lower insets in G, J), 10 μm (remaining panels). (A-T), single confocal z-sections. |
|
Morphological rescue of some moe- embryonic defects by injection of the epb4.1l5 constructs. (A) In wild-type embryos at 60 hpf, brain ventricles are visible, and the RPE is uniform (inset). (B) In moe- embryos, brain ventricles are reduced in size or absent, pericaridal edema is pronounced, and the RPE is patchy (inset). (C) In moe- embryos injected with epb4.1l5short+long_PBD mRNA, brain ventricles are reduced in size or absent, pericaridal edema is pronounced, but the RPE is normal (inset). (D) moe- embryos injected with epb4.1l5long+short_PBD mRNA, brain ventricles are absent or absent and pericardial edema is pronounced, and RPE defects are milder than those in uninjected moe- embryos. (E) A magnified view of the RPE of a 60 hpf wild-type embryo shows that it is uniform and the cells are confluent. (F) In a wild-typounced, and RPE defects are milder than those in uninjected moe- embryos. (E) A magnified view of the Re retina at 4 dpf, GFP+ rods localize next to the RPE and lamination is apparent. In 60 hpf moe- mutants, the integrity of the RPE varies from mild (G), to moderate (I), to severe (H). However, GFP+ rods are mislocalized in all moe- mutants 4 dpf (H, J, L). The integrity of the RPE is improved and nearly normal in a 60 hpf moe- mutant injected with epb4.1l5long+short_PBD mRNA (M) and most GFP+ rods are adjacent to the RPE (N). (O) A wild-type embryo injected with epb4.1l5long+short_PBD showing brain ventricles that are reduced or absent. (P) At 30 hpf Epb4.1l5long+short_PBD is cortically localized, upper inset is a 2x magnification of Epb4.1l5long+short_PBD localization. Scale bars, 10 μm (F), 50 μm (lower insets in F). |
|
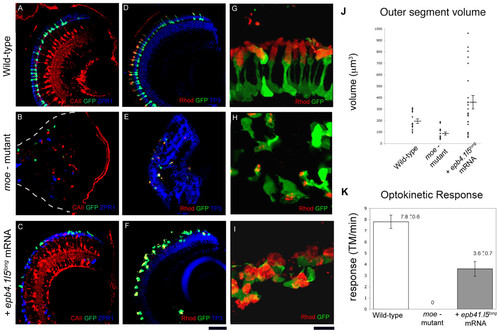
Injection of epb4.1l5long mRNA into moe- mutants restores retinal lamination but not normal photoreceptor morphology at 6 dpf. In wild-type (A) and epb4.1l5long mRNA injected moe- mutants (C), Müller glial cells are radially oriented (labeled with anti-Carbonic Anhydrase II, red), and GFP+ rods and double cones (ZPR-1 antibody, blue) are adjacent to the RPE. (B) In moe- mutants, Müller glial cells (red), and rods (green) and double cones (blue) are fewer in number and are localized ectopically. White dashed line indicates the back of the eye. (D) In wild-type larvae, distinct nuclear layers (labeled by TO-Pro3, blue) are visible and outer segments labeled by anti-Rhodopsin (red) are aligned radially and project toward the RPE. (E) In moe- mutants, distinct nuclear layers are absent and rod morphology is abnormal. (F) In epb4.1l5long mRNA injected moe- mutants, distinct nuclear layers are visible but rod photoreceptor morphology is abnormal. (G, H, I) High magnification confocal z-projections of GFP+ rods labeled with anti-Rhodopsin (red) in wild-type (G), moe- mutant (H), and epb4.1l5long mRNA injected moe- mutant (I). (J) Outer segment volume at 6 dpf in wildtypes, moe- mutants, and epb4.1l5long mRNA injected moe- mutants. The average volume of rod outer segments in epb4.1l5long mRNA injected moe- mutants is 362 μm3 (+/- 57.1) which was significantly greater than the wild-type volume of 197 μm3 (+/- 18.5), p = 0.025. The volume of moe- rod outer segments was 90.9 μm3 (+/- 12.3), significantly smaller than wild-type rods (p < 0.001). Significance was determined using the Students T-test. (K) Optokinetic response was measured at 5 dpf in wild-types, moe- mutants and epb4.1l5long mRNA injected moe- mutants. Wild-type larvae exhibited an average of 7.8 (+/- 0.6) tracked movements (TM)/minute, epb4.1l5long mRNA injected moe- mutants exhibited a reduced OKR (3.6 (+/- 0.7) TM/minute). moe- mutants exhibited no OKR. N = 10 for each group. Scale bars, 40 μm (A-F), 10 μm (G-I). (A-F, single confocal z-sections, G-I, z-projections). |
|
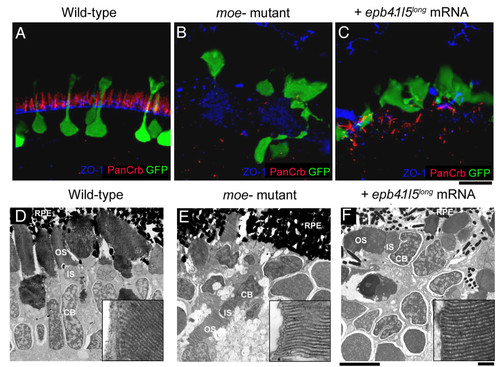
The outer limiting membrane (OLM) is not restored in moe- mutants by injection of epb4.1l5long mRNA. (A) In wild-type larvae at 6 dpf, Crumbs proteins localize just apical to the OLM, which is labeled by anti-ZO-1. (B) In moe- mutants, very little Crumbs protein is visible and ZO-1 labeling is disorganized. (C) In epb4.1l5long mRNA injected moe- mutants, streaks of panCrb labeling are visible, but an organized OLM is absent. Ultrastructural transmission electron microscopic analysis at 6 dpf in wild-type (D), moe- mutants (E) and epb4.1l5long mRNA injected moe- mutants (F) retinas. Electron dense outer segments are seen in all individuals. Insets, higher magnifications of rod outer segments showing regular disc stacking is present in all individuals (100,000X). RPE, retinal pigmented epithelium; OS, outer segments; IS, inner Segments; CB, cell body. (A-C) are confocal z-projections, scale bar 10 μm (D-F). Scale bars, 5 μm (D-F), 100 nm (insets). |