- Title
-
Mammalian germ-line transgenesis by transposition
- Authors
- Dupuy, A.J., Clark, K., Carlson, C.M., Fritz, S., Davidson, A.E., Markley, K.M., Finley, K., Fletcher, C.F., Ekker, S.C., Hackett, P.B., Horn, S., and Largaespada, D.A.
- Source
- Full text @ Proc. Natl. Acad. Sci. USA
|
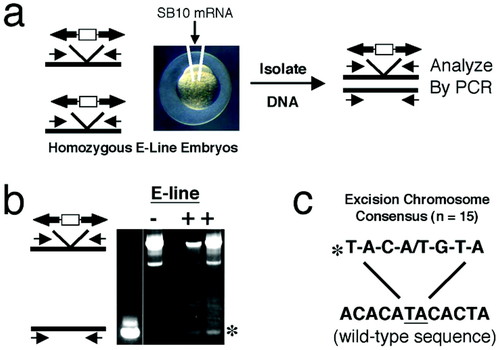
Verifying activity of SB10 mRNA. In vitro-transcribed SB10 mRNA was injected into zebrafish embryos homozygous for a transposon insertion (line E). Embryos are then collected after 24 h, and genomic DNA is extracted from them (a). PCR is then performed by using primers that flank the site of transposon insertion. Transposition can then be detected by appearance of the PCR product from the transposon insertion site after excision (b). Excision can be confirmed by sequencing the PCR product to detect the three nucleotide footprint (c). The sequence of the insertion site following transposon excision is indicated (*). Two different base-pair footprints are found in roughly equal proportion, CAG or CTG. |
|
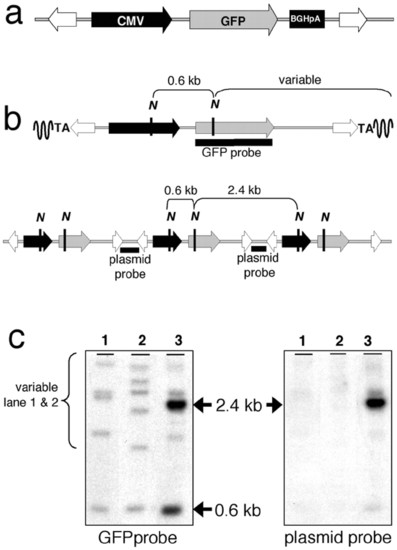
Structure and analysis of transposon integration sites. (a) The structure of the linear transposon that was used for pronuclear injection is shown. The transposon contains a GFP reporter (gray arrow) driven by the immediate early CMV promoter (black arrow) followed by the bovine GH polyadenylation site (black box) and flanked by the IR/DRs (white arrows). (b Upper) The structure of a Sleeping Beauty-mediated integration site including the expected fragments from a NcoI (N) digest. (b Lower) The structure of a random integration site and NcoI restriction map. The probes used for Southern blot analysis are also defined as black bars. (c) Examples of three embryonic DNAs analyzed by Southern blotting are shown. The results shown were achieved with the GFP probe (Left) and the plasmid probe (Right). |