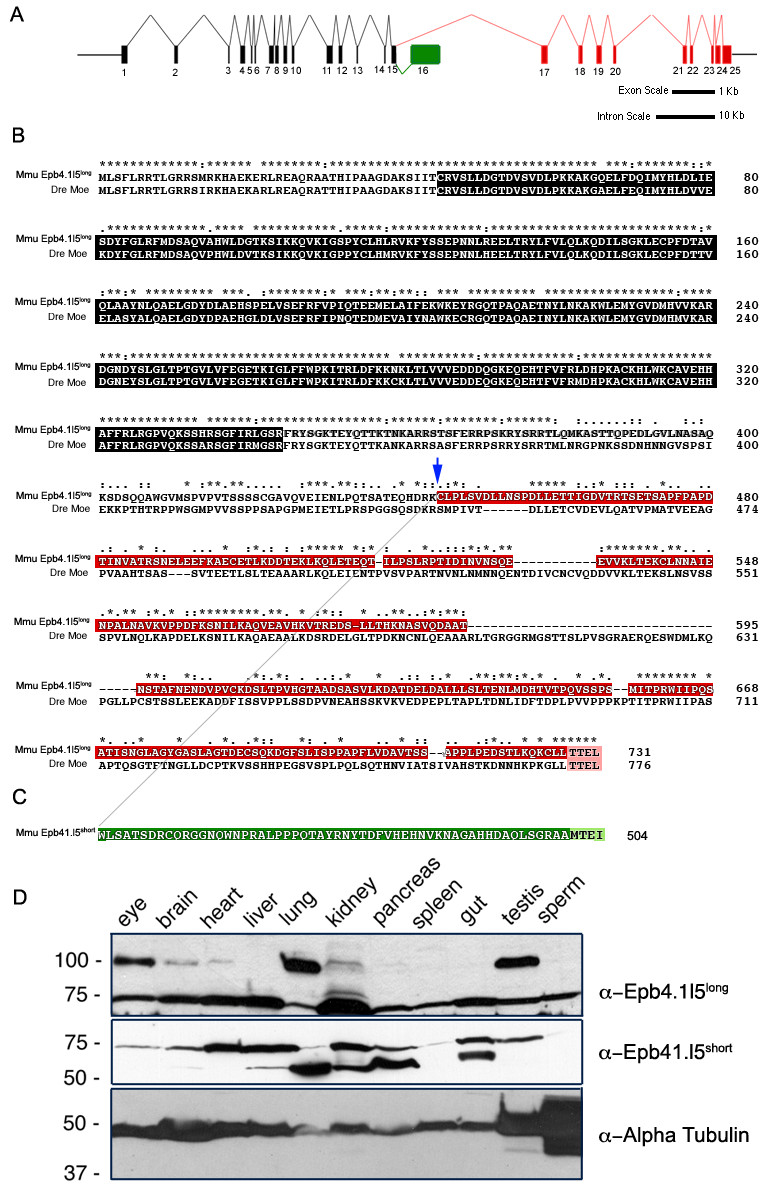
Fig. 2 Genomic structure of the mouse epb4.1l5 locus and expression of its two major splice isoforms. (A) Diagram of the inton/exon structure of the Mus musculus epb4.1l5 locus. Exons that are common to both isoforms are black, the exons unique to epb4.1l5long are indicated in red and the unique exon in epb4.1l5short is indicated in green. Bars represent 1 kb and 10 kb scales for exon and intron lengths, respectively. (B) ClustalX alignment of mouse Epb4.1l5long and zebrafish Moe. Ymo1long and Moe share a high degree of homology within the FERM domain (black). Ymo1long and Ymo1short are identical up to Lysine 444 (blue arrow) and then alternately spliced into the long (red) and short (green) isoforms. Moe and Epb4.1l5long, and Epb4.1l5short have predicted C′terminal PDZ-binding domains (Pink [TTEL]) and (light green [MTEI]). (*) identical, (:) highly conserved, (.) Moe and Epb4.1l5long, and Epb4.1l5short have predicted C′terminal PDZ-binding domains (Pink moderately conserved. (C) Western analysis of mouse tissues with antibodies raised against the unique C′terminal sequences of Epb4.1l5long and Epb4.1l5short. Two bands are immunoreactive with the anti-Epb4.1l5long antibody, one migrates at the expected molecular weight of Epb4.1l5long (100 kDa) and is present in the eye, brain, heart, lung, kidney, and testis. An additional band migrates at 75 kDa, which is probably non-specific. Two bands are recognized by the affinity purified Epb4.1l5short antibody. A lower band migrates at the predicted molecular weight of 56 kDa and is present in brain, liver, lung, kidney, pancreas, and gut. A second band with a broad expression pattern migrates at approximately 75 kDa. Anti-α-Tubulin was used as a loading control.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ BMC Dev. Biol.